Core
Gene & accession numbers
Introduction
Results
Screening transcriptomes of different leaves identified a likely trichome-regulatory MYB TF gene
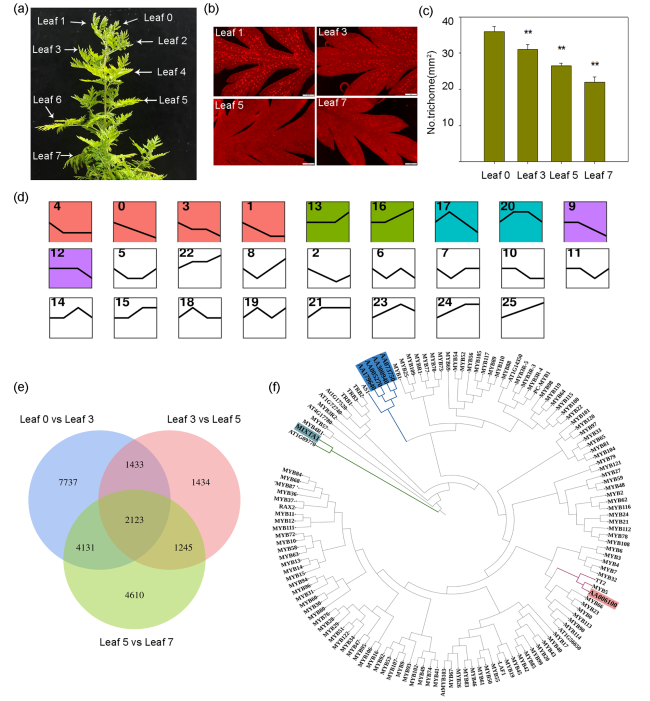
Fig. 1 Transcriptome sequencing and differential gene analysis of leaves 0, 3, 5 and 7 of A. annua plants. a Leaves from a three-month-old plant, numbered from top (youngest) to bottom (oldest). b Glandular trichomes on leaf 0, leaf 3, leaf 5 and leaf 7 of A. annua visualized by fluorescence microscopy. Autofluorescence of glandular trichomes was captured with λex = 480 nm and λem = 535 nm. c Trichome density in A. annua leaf 0, leaf 3, leaf 5 and leaf 7. Asterisks indicate significant differences relative to leaf 0 (Student’s t-test; **, P < 0.01). Data are means ± standard error (SE). d Clustering analysis of 2,123 genes in different leaves. The x axis represents leave samples from four developmental stages (leaf 0, leaf 3, leaf 5 and leaf 7); the y axis represents gene expression. Twenty-five clusters were identified. e Venn diagram showing the extent of overlap between differentially expressed genes in the comparisons leaf 0 vs leaf 3; leaf 3 vs leaf 5; and leaf 5 vs leaf 7. f Maximum likelihood phylogenetic tree reconstructed based on six identified A. annua transcription factors and 131 Arabidopsis thaliana transcription factors |
Characterization of TLR3 expression
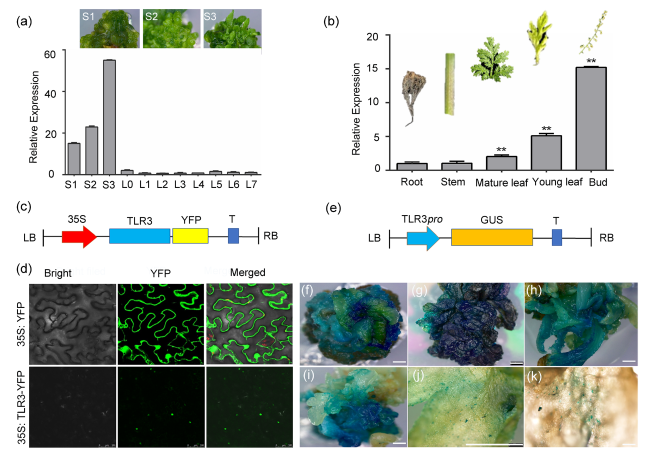
Fig. 2 TLR3 expression level and TLR3 subcellular localization in A. annua. a Relative TLR3 expression level in callus (stages S1-S3) and leaves 0-7 (L1-L7). S1, callus primary stage; S2, callus middle stage; S3, shooting stage. b Relative TLR3 expression level in roots, stems, old leaves, young leaves and flowers. c Schematic of the TLR3-YFP construct, driven by the 35S promoter. YFP, yellow fluorescent protein; LB, left border; RB, right border; T, termination sequence. d TLR3 is localized in the nucleus in Nicotiana benthamiana leaf epidermal cells. Free YFP (35S:YFP) was used as a control. e Schematic of the TLR3pro:GUS reporter construct. GUS, ß-glucuronidase. f-k GUS staining of A. annua callus and tissues from TLR3pro:GUS transgenic materials. Scale bars, 1 mm |
A. annua TLR3 negatively regulates trichome density and branching in Arabidopsis
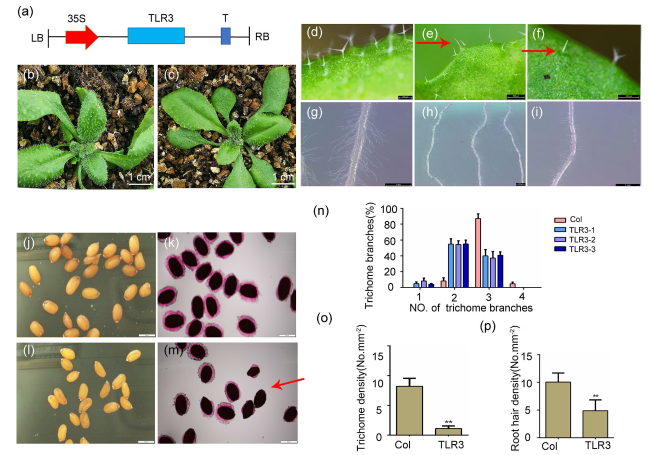
Fig. 3 Heterologous expression of TLR3 in Arabidopsis thaliana represses trichome development. a Schematic of the 35S:TLR3-PHB construct. PHB is the vector used for gene overexpression in plants. b Representative nontransgenic Col-0 plant. c Representative TLR3-OE plant. d-n Transgenic plants show fewer trichomes, and trichomes have fewer branches (e, f, n). Red arrows point to a trichome with a single branch (e) and a trichome with two branches (f). As compared to Col-0 (g), the transgenic plants have a lower density of root hairs (h, i). Seeds of Col-0 and transgenic plants, without (j, l) and with (k, m) ruthenium red staining. The transgenic seeds are abnormal seeds, with a defect in mucilage (stained by ruthenium red; m). n Distribution of the number of trichome branches. o Distribution of the number of trichome density. p Distribution of the number of root hair density. Asterisks indicate significant differences relative to Col-0 (Student’s t-test; **, P < 0.01). Data are means ± SE |
TLR3 physically interacts with NFY1 and CycTL
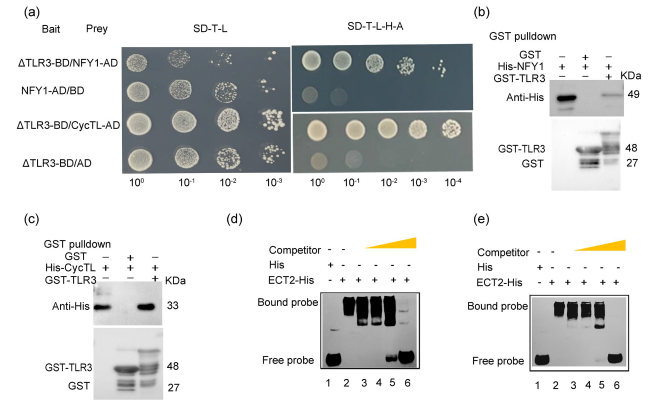
Fig. 4 TLR3 can interact with NFY1 and CycTL in yeast two-hybrid and pull-down assays and can bind to the UGUAA, UGUAU and GGACU of ECT2 mRNA. a Yeast two-hybrid (Y2H) assay indicating that truncated TLR3 (∆TLR3, amino acids [aa] 1-111) interacts with NFY1 and CycTL. The bait and prey plasmids were co-transformed into yeast and positive colonies were selected on synthetic defined (SD) medium lacking Trp, Leu, His and Ade. b-d Pull-down assay indicating that AaTLR3 interacts with NFY1 (b), CycTL (c). Recombinant GST-TLR3, His-NFY1 and His-CycTL were produced in E. coli Rosetta (DE3) and purified. After incubation in vitro, the protein mixture was separated by SDS-PAGE gel. Proteins were detected with anti-His and anti-GST antibodies. d, e ECT2-His interacts with the UGUAA, UGUAU and GGACU of TLR3 mRNA on RNA-EMSA. Recombinant ECT2-His was produced in E. coli Rosetta (DE3) and purified. The concentration ratios of free probe and bound probe in the third to sixth lane were 1:1, 1:10, 1:100 and 1:1000, respectively |
TLR3 regulates trichome branching
TLR3 negatively modulates trichome development and artemisinin yield in A. annua
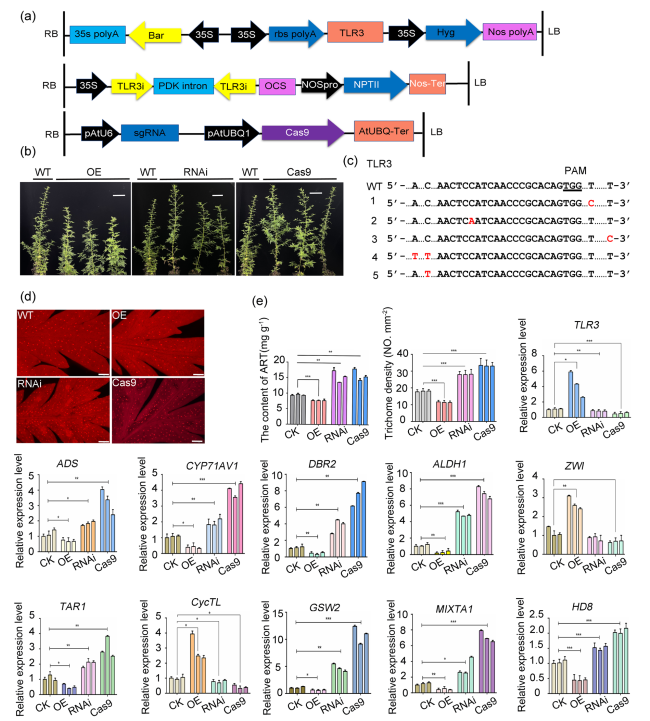
Fig. 5 Phenotypes and relative expression of genes in transgenic A. annua plants. a Schematics of the vectors PHB-TLR3, RNAi-TLR3 and Cas9-TLR3. b Comparison of plant height in WT and TLR3 transgenic plants. Scale bars, 10 cm. c Identification of TLR3 mutations in TLR3-Cas9 plants. The bases in red are insertions generated by genome editing. d Trichome density on the leaves of WT, TLR3-OE, TLR3-RNAi and TLR3-Cas9 plants as visualized by fluorescence microscopy. Scale bars, 100 µm. e Content of ART (artemisinin), trichome density and relative expression levels of the indicated genes in transgenic TLR3 A. annua plants |
Overexpression of TLR3 leads to abnormal trichomes in A. annua
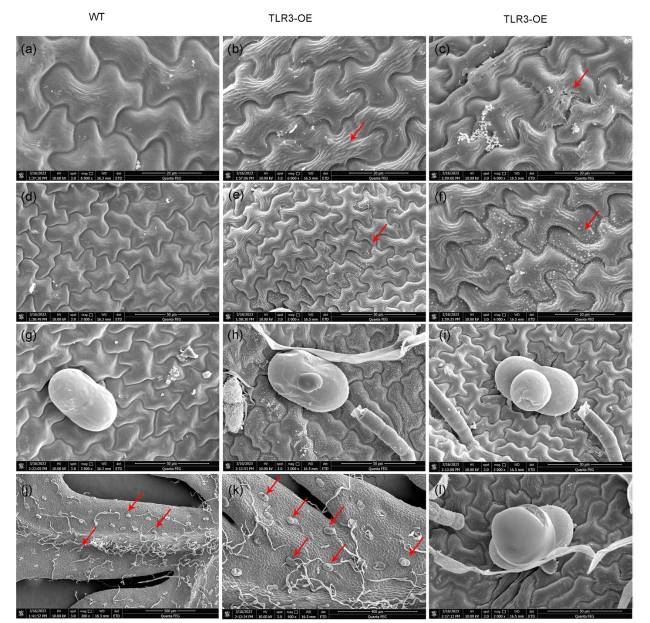
Fig. 6 Scanning electron microscopy (SEM) analysis of the cuticle and trichomes on the surface of A. annua leaves. a Smooth surface of WT A. annua leaves. b Coarse and fibrous surface of A. annua TLR3-OE leaves. c A TLR3-OE leaf with detached cuticular wax visible on its surface. d Smooth surface of WT A. annua leaves without white wax crystals. e, f TLR3-OE leaves show uneven distribution of white wax crystals. g A normal glandular trichome on a WT A. annua leaf. (h, i, l) Abnormal glandular trichome on a leaf of an A. annua TLR3-OE plant. j Few abnormal glandular trichomes (red arrows) are visible on WT A. annua leaves. k Many abnormal glandular trichomes (red arrows) are visible on the leaves of A. annua TLR3-OE plants |
Discussion Overexpression of TLR3 alters cuticle biosynthesis in A. annua
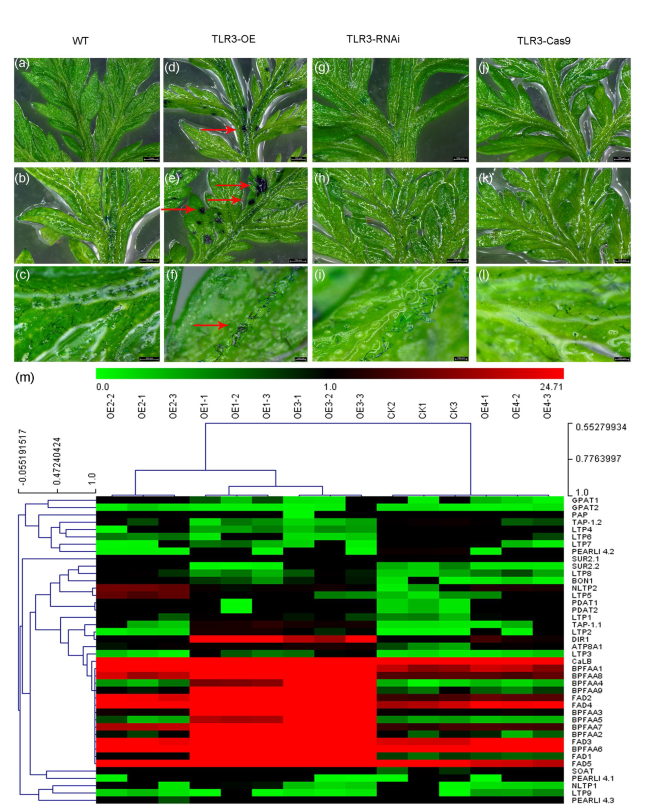
Fig. 7 TLR3 is involved in the biosynthesis of cuticle in A. annua. a-l Young leaves of WT (a), TLR3-OE (d), TLR3-RNAi (g) and TLR3-Cas9 (j) plants treated with toluidine blue (TB); mature leaves of WT (b), TLR3-OE (e), TLR3-RNAi (h) and TLR3-Cas9 (k) plants treated with TB; and leaf veins of WT (c), TLR3-OE (f), TLR3-RNAi (i) and TLR3-Cas9 (l) plants were treated with TB. Red arrows point to sites that took up toluidine blue. m Hierarchical clustering analysis of genes involved in biosynthesis of lipids in WT A. annua and TLR3-OE leaves. The color scale at the top represents the log-transformed reads per kilobase per million mapped reads |
Discussion
TLR3 negatively regulates biomass by interacting with NFY1
TLR3 negatively regulates the density and branching of trichomes
TLR3 modulates cuticle biosynthesis in A. annua
Mode of trichome and cuticle regulation by TLR3
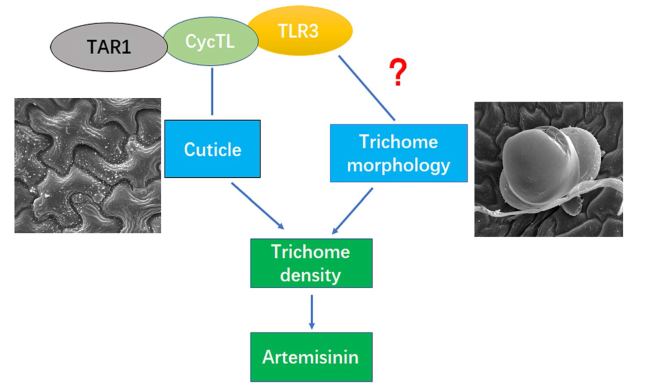
Fig. 8 Model for TLR3 regulation of cuticle and trichome morphology. TLR3 interacts with CycTL, which interacts with TAR1. The three proteins may regulate the distribution of cuticle on A. annua leaves. TLR3 also affects trichome morphology, resulting, for instance, in club trichomes, calabash trichomes. During trichome development, genes influence cuticle biosynthesis and trichome morphology, altering the initial and phenotype of trichome in A. annua |