Core
Gene & Accession Numbers
Introduction
Results
ATAC-seq identified and predicted floral organ identity gene in P. equestris var.trilip
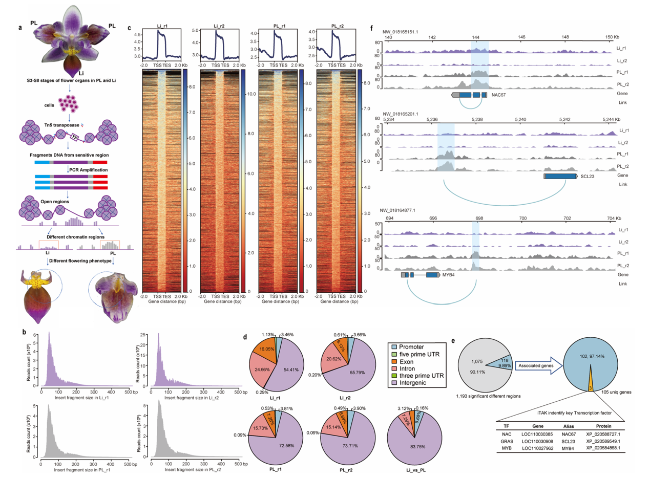
Fig. 1 ATAC-seq is used to identify key transcription factors in lip development of P. equestris var.trilip. a Diagram of P. equestris var.trilip flower with petal (PL) and lip (Li) phenotypes, and method of using ATAC-seq to identify accessible regions in the genome of both mutants. b Distribution of insert fragment size in PL and Li samples. c Distribution of peaks within the 2-kb interval upstream and downstream of the transcription start site (TSS). d Proportions of genome regions covered by fragments generated in ATAC-seq analysis in PL samples, Li samples, and Li vs. PL. e Significantly different regions between PL and Li samples, and transcription factor (TF) genes among differentially expressed genes (DEGs). f Differences in transcription of three TF genes between PL and Li samples |
Co-expression of PeNAC67 and PeSCL23 with B-class MADS-box genes during flower development
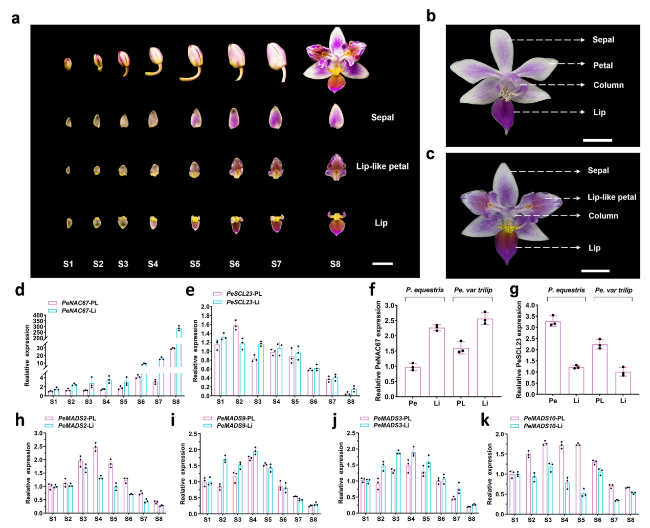
Fig. 2 Expression of TF genes in P. equestris var.trilip and P. equestris flowers. a PL and Li from S3-S8 development flowers in the P. equestris var.trilip. Scale bar = 1 cm. b and (c) Different flower organs of P. equestris and P. equestris var.trilip. Scale bar = 1 cm. d, (e) and (h-k) Relative transcript levels of PeNAC67 (d), PeSCL23 (e), PeMADS2 (h), PeMADS3 (i), PeMADS9 (j) and PeMADS10 (k) in different organs of P. equestris var.trilip at different developmental stages. f and (g) Relative transcript levels of PeNAC67 (f), PeSCL23 (g) in the petal and lip of P. equestris and relative transcript levels in the lip-like petal and lip of P. equestris var.trilip at S1-S8 development stage. The expression patterns of all genes were determined using three replicates and were normalized using PeActin4. OAP3-1: PeMADS2, OAP3-2: PeMADS3, OAGL6-2: PeMADS9, OAGL6-1: PeMAD10 |
PeNAC67 correlating with MADS3 affected petal specialization in P. equestris var.trilip
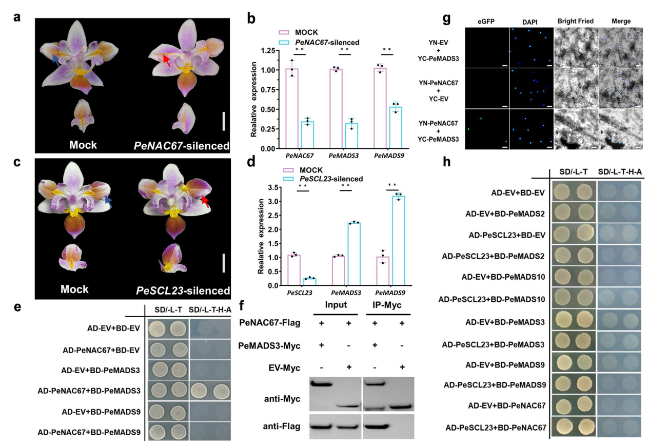
Fig. 3 Phenotypes of PeNAC67 and PeSCL23 silencing lines and interaction between PeNAC67 with PeMADS3. a and (c) Mock and VIGS-treated lines with silenced expression of PeNAC67 (a) and PeSCL23 (c). The red arrows in (a) and (c) represent the part where the mutation occurs after VIGS treatment, and the blue arrows in (a) and (c) represent the corresponding part in the Mock. b and (d) Relative transcript levels of PeNAC67, PeMADS3 and PeMADS9 in Mock and VIGS-treated lines (b) and relative transcript levels of PeSCL23, PeMADS3 and PeMADS9 in Mock and VIGS-treated lines (d), asterisk in (b) and (d) indicated significant differences compared with the control, with one asterisk indicating P < 0.05 and two asterisks indicating P < 0.01. e Yeast two hybrid assay between PeNAC67 and PeMADS3/PeMADS9. f Co-IP was used to detect the interaction between PeNAC67-Flag and PeMADS3-Myc proteins transiently expressed in tobacco leaves. g Bimolecular fluorescence complementation (BiFC) assay was introduced to detect the interaction between PeNAC67 and PeMADS3 in tobacco leaves with DAPI staining. h Yeast two hybrid assay of PeSCL23 with B-class MADS-box protein |
PeKAN2 as the key factor in conjugate regulation between PeNAC67 and PeSCL23 during specialization process from petal to lip
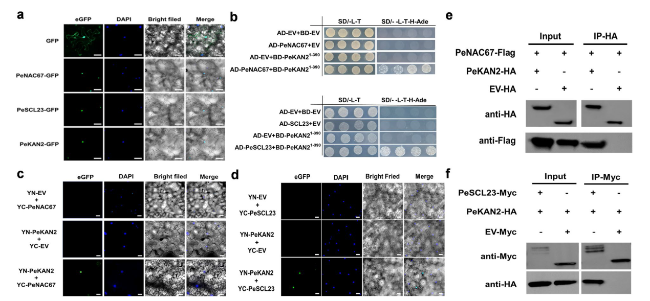
Fig. 4 Analyses of interactions of PeNAC67 and PeSCL23 with PeKAN2. a Subcellular localization of PeNAC67-GFP, PeSCL23-GFP and PeKAN2-GFP proteins in Nicotiana benthamiana leaf epidermal cells with DAPI staining. Scale bars = 50 μm. b Yeast two hybrid assay of PeNAC67 and PeSCL23 with PeKAN2(1-390). c and (d) BiFC assay was introduced to detect the interaction PeNAC67 and PeSCL23 with PeKAN2 in tobacco leaves. e and (f) Co-IP was used to detect the interaction between PeNAC67-Flag and PeSCL23-Myc with PeKAN2-HA proteins transiently expressed in tobacco leaves |
Transcript level of PeKAN2 was correlated with those of PeMADS3/PeMADS9 and together regulate specialization process from petal to lip
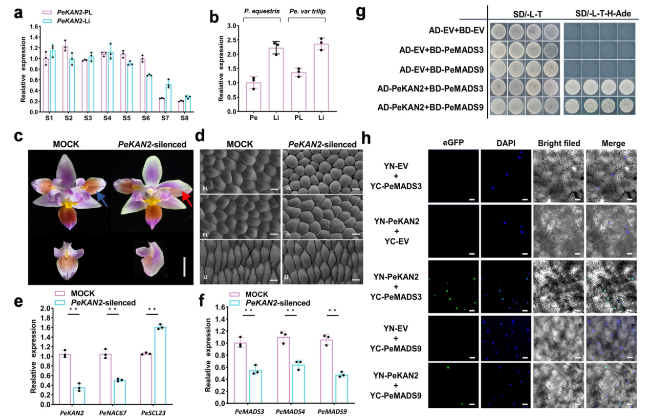
Fig. 5 PeKAN2 expression during flower development and analyses of the role of PeKAN2. a Relative transcript levels of PeKAN2 in different organs of P. equestris var.trilip at different developmental stages(from stages S1 to S8 development). b Relative transcript levels of PeKAN2 in the petal and lip of P. equestris and relative transcript levels in the lip-like petal and lip of P. equestris var.trilip at S8 stage. c The phenotypes of PeKAN2-silenced lines. The red arrow represents the part where the mutation occurs after VIGS treatment, and the blue arrow represents the corresponding part in the Mock. d The epidermal cells of wild-type control (Left) and those of PeKAN2-silenced (right) flowers. Bar = 50 μm. e Relative transcript levels of PeKAN2, PeNAC67 and PeSCL23 in Mock and VIGS-treated lines. f Relative transcript levels of PeMADS3, PeMADS4 and PeMADS9 in Mock and VIGS-treated lines. g Yeast two hybrid assay of PeMADS3/PeMADS9 with PeKAN2. h BiFC assay was introduced to detect the interaction PeMADS3/PeMADS9 with PeKAN2 in tobacco leaves. Values are means ± SDs (n = 3). Asterisks in e and f indicate significant differences compared with the control, with one asterisk indicating P < 0.05 and two asterisks indicating P < 0.01 |
Interaction between PeKAN2 and PeMADS3 was enhanced by PeNAC67 but inhibited by PeSCL23
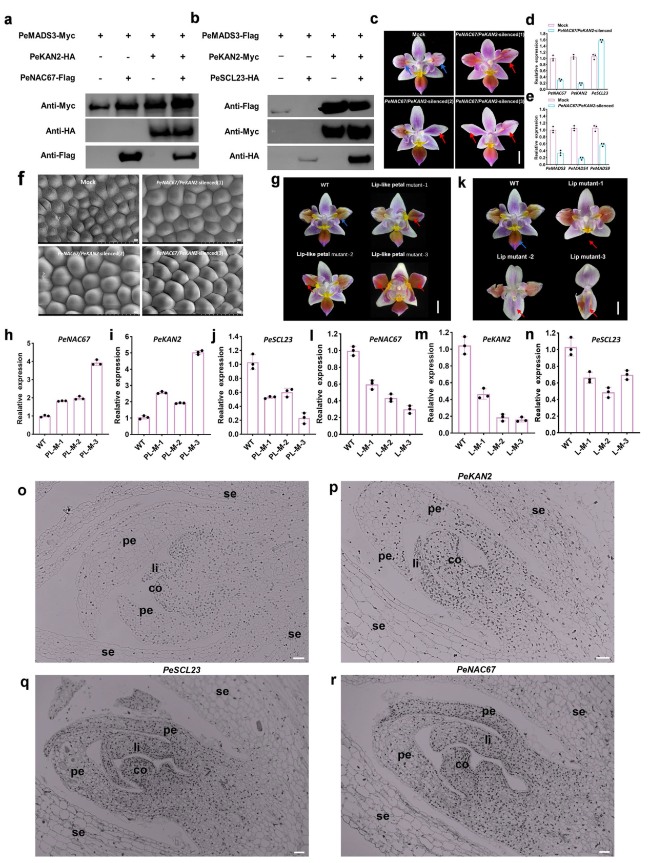
Fig. 6 Biological function analysis of PeNAC67, PeKAN2 and PeSCL23 in Phalaenopsis. a The fusion constructs of PeNAC67-Flag, PeMADS3-Myc, PeKAN2-HA tags were transformed into P.Big chill in different combinations, and Myc/Flag/HA antibody was used for immunoprecipitation. b The fusion constructs of PeSCL23-HA, PeMADS3-Flag and PeKAN2-Myc tags were transformed into P.Big chill in different combinations, and Myc/Flag/HA antibody was used for immunoprecipitation. c Representative phenotypic analysis for three independent PeNAC67 and PeKAN2 double silencing lines are presented. The red arrow represents the part where the mutation occurs after PeNAC67 and PeKAN2 VIGS treatment, and the blue arrow represents the corresponding part in the Mock. d and (e) Relative transcript levels of PeKAN2, PeNAC67 and PeSCL23 in Mock and VIGS-treated lines (d) and relative transcript levels of PeMADS3, PeMADS4 and PeMADS9 in Mock and VIGS-treated lines (e). f The epidermal cells of wild-type control and those of PeNAC67/PeKAN2 silenced flowers representative phenotypic analysis for three independent silencing lines was presented. Bar = 50 μm. g “Lip-like petal” mutant phenotype of P. equestris var.trilip. The red arrow represents the part where the mutation of “Lip-like petal”. h-j The expression of PeNAC67, PeKAN2 and PeSCL23 in “Lip-like petal” mutant phenotype of P. equestris var.trilip. (k) “Lip” mutant phenotype of P. equestris var.trilip. The red arrow represents the part where the mutation of “Lip petal”. l-n The expression of PeNAC67, PeKAN2 and PeSCL23 in lip mutant phenotype of P. equestris var.trilip. o-r In-situ localization of PeKAN2 (p), PeSCL23 (q) and PeNAC67 (r) transcripts in P. equestris flower buds. Longitudinal sections were hybridized with DIG-labeled antisense. o A negative control was performed by sense probe. co, column; pe, petal; se, sepal; li, lip. Bar = 50 μm |
Discussion
The first successful application of ATAC-Seq to identify important genes in a horticultural flower
New function of non-MADS genes in flower organ development
PeKAN2 regulated lip development by serving as a bridge between PeNAC67 and PeSCL23
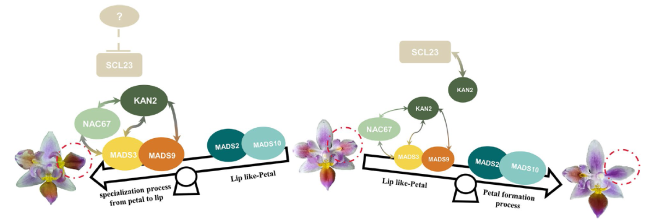
Fig. 7 Model of PeNAC67, PeSCL23, PeKAN2 cooperation with MADS box protein to regulate the lip development in P. equestris var.trilip. During the specialization process from petal to lip, PeSCL23 translation was inhibited, PeNAC67 interacted with PeKAN2, and enhanced the stability of PeMADS3; Meanwhile PeKAN2 also could correlating with PeMADS9. The development of the lip was promoted through enhancing PeMADS3 activity. During the formation process of petals, PeSCL23 and PeNAC67 competitively interacted with PeKAN2, which brought about the decreasing of the PeMADS3 activity, and instead promoting petal formation |