Core
Gene & accession numbers
Introduction
Results
Identification of SSPs in tomato
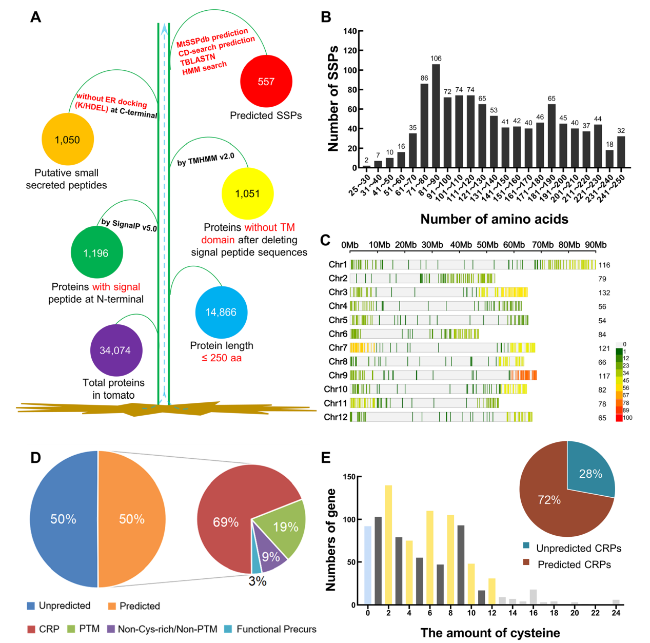
Fig. 1 Identification and classification of SSPs in tomato. A The procedure of tomato SSP identification. B The number of SlSSPs with different protein length ranges. C Chromosomal distribution of SlSSPs. SlSSP density plot on each chromosome represented by number of SSPs within 1 Mb window size. Chromosome distribution visualization was created by an online platform (https://www.bioinformatics.com.cn). D The percentage of different types of SlSSPs predicted in MtSSPdb (https://mtsspdb.noble.org/database/). CRP, cysteine-rich peptide; PTM, post-translational modified. E The prediction of CRPs in tomato. The column represents amount of SlSSPs with different number of cysteines after deleting signal peptide sequences, the pie represents the percentage of predicted CRPs and unpredicted CRPs in MtSSPdb |
Classification of SSPs in tomato
Table 1 Predicted SSP families in tomato |
| Class | SSP family | Description | Mode of action | Number of peptides |
|---|---|---|---|---|
| Post-translationally modified (PTM) | CEP | C-terminally encoded peptide | Signal | 21 |
| CLE | Clavata/Embryo Surrounding Region | Signal | 43 | |
| GLV/RGF/CLEL | Golven/Root Growth Factor | Signal | 12 | |
| IDA | Inflorescence Deficient in Abscission | Signal | 8 | |
| PIP | PAMP-induced Secreted Peptide | Signal | 4 | |
| PSK | Phytosulfokine | Signal | 8 | |
| PSY | Plant Peptide Containing Sulfated Tyrosine | Signal | 11 | |
| Cysteine rich | 2SA | 2S Albumin | Antimicrobial | 2 |
| ECL | Egg Cell 1-Like | Signal | 10 | |
| EPFL | Epidermal Patterning Factor-Like | Signal | 12 | |
| GASA | Gibberellic Acid Stimulated in Arabidopsis | Signal | 20 | |
| HEVEIN | Hevein | Antimicrobial | 9 | |
| Kunitz | Kunitz-P trypsin inhibitor | Peptidase inhibitor | 17 | |
| LAT52-POE | LAT52/Pollen Ole e 1 Allergen | Signal | 19 | |
| MEG | Maternally Expressed Gene | Signal | 2 | |
| N26 | Nodulin26 | Signal | 1 | |
| nsLTP | non-specific Lipid Transfer Protein | Signal | 122 | |
| PCY | Plantcyanin/Chemocyanin | Signal | 46 | |
| Plant Defensin | Antimicrobial | 51 | ||
| RALF | Rapid Alkalinization Factor | Signal | 11 | |
| RC | Root Cap | Signal | 2 | |
| STIG-GRI | Stigma1/GRI | Signal | 10 | |
| T2SPI | Potato type II proteinase inhibitor | Peptidase inhibitor | 13 | |
| THL | Thionin-like | Antimicrobial | 18 | |
| TPD | Tapetum Determinant 1 | Signal | 6 | |
| Kaz | Kazal family inhibitors | Peptidase inhibitor | 2 | |
| PDL | Plant Defensin-like | Antimicrobial | 2 | |
| LCR | Low-molecular weight Cys-rich | Unknown | 3 | |
| TAX | Taximin | Signal | 3 | |
| SCR/SP11 | S-locus Cysteine Rich | Signal | 4 | |
| Non-Cys-rich/Non-PTM | CTLA | Cytotoxic T-lymphocyte antigen-2 alpha | Peptidase inhibitor | 5 |
| GRP | Glycine-rich Protein | Unknown | 6 | |
| PhyCys | Phytocystatin | Peptidase inhibitor | 9 | |
| PNP | Plant Natriuretic Peptide | Signal | 7 | |
| PRP669 | Pro-rich Protein Group 669 | Unknown | 4 | |
| Subln | Subtilisin inhibitor | Peptidase inhibitor | 17 | |
| Functional Precursor | CAPE | CAP-derived Peptide | Signal | 14 |
| MtSUBPEP | Subtilisin-embedded Plant Elicitor Peptide | Signal | 3 | |
| Total | 557 |
Fig. 2 Structural characteristics of proproteins of typical SlSSPs. A The multiple sequence alignment of typical SSP families. CLE represents PTM peptides, RALF represents CRPs, PNP represents Non-Cys-rich/No-PTM peptides. B, C Two putative new SSP families predicted by manual analysis. Red single lines indicate signal peptides, and red double lines indicate putative mature peptides. Yellow arrows indicate the putative endoprotease dibasic cleavage sites (RR), and blue arrows indicate the conserved tyrosine residues. Gray underlines indicate the “K(V/I)(V/I)D” domain and red stars indicate the conserved “LS” and “IA” domains in (A). Black stars indicate the conserved cysteine residues. Brackets represent mature peptide sequence modules of SSP families from (B, C). Superscript and subscript numbers represent position of cysteine residues and the number of amino acids, respectively.“n” indicates any number |
Identification of unannotated small secreted peptides in tomato
Table 2 Bioinformatic identification and filtering of tomato peptide-encoding sORFs |
| Tomato Chromosome | No. sORFs kept after each sequential filter | ||||
|---|---|---|---|---|---|
| 25-250 aa | N-terminal SP | Non-TM | Putative SSP | Known SSP families | |
| 0 | 48,555 | 590 | 570 | 570 | 0 |
| 1 | 362,481 | 7,262 | 7,064 | 7,064 | 9 |
| 2 | 196,102 | 3,884 | 3,796 | 3,794 | 6 |
| 3 | 251,100 | 4,800 | 4,668 | 4,668 | 6 |
| 4 | 258,114 | 5,108 | 4,998 | 4,998 | 2 |
| 5 | 274,533 | 5,583 | 5,433 | 5,432 | 3 |
| 6 | 179,709 | 3,503 | 3,410 | 3,409 | 2 |
| 7 | 279,966 | 5,515 | 5,383 | 5,382 | 8 |
| 8 | 263,811 | 5,203 | 5,076 | 5,076 | 6 |
| 9 | 287,462 | 5,986 | 5,840 | 5,840 | 3 |
| 10 | 273,707 | 5,573 | 5,447 | 5,447 | 2 |
| 11 | 217,273 | 4,320 | 4,200 | 4,200 | 6 |
| 12 | 282,705 | 5,555 | 5,426 | 5,426 | 6 |
| Total | 3,175,518 | 62,882 | 61,311 | 61,306 | 59 |
sORFs small ORFs, SP Signal peptide, TM Transmembrane |
Expression patterns of SlSSPs in different tissues and under drought stress
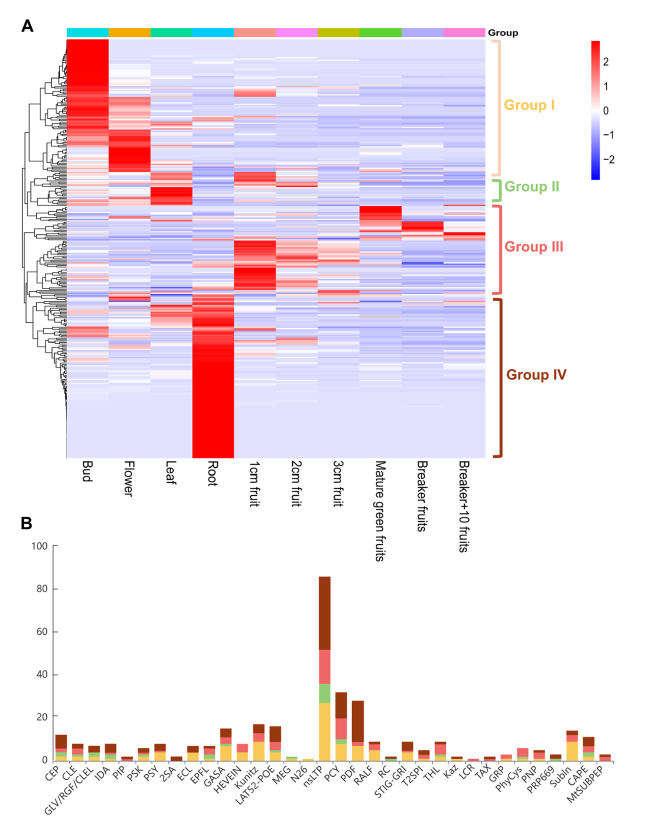
Fig. 3 The expression pattern of tomato SSP genes in different tissues. A Heatmap visualization of the expression pattern of tomato SSP genes in bud, flower, leaf, root, 1 cm fruit, 2 cm fruit, 3 cm fruit, mature green fruits, breaker fruits and breaker + 10 fruits. The expression data was downloaded from Tomato Expression Database (D004, http://ted.bti.cornell.edu/). The heatmap was generated by TBtools. B The number of SSP members of different families in group I (yellow), group II (green), group III (red), and group IV (brown) |
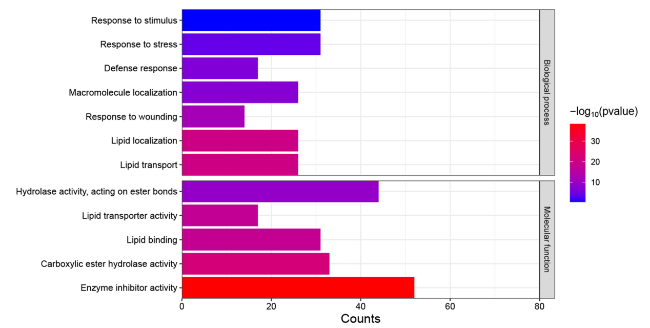
Fig. 4 GO analysis of tomato SSPs. Go terms involved in stress response. X-axis represents the number of genes in each GO Term, Y-axis is GO Terms. The color indicates P value. Go analysis was done by AgriGO v2.0 (http://systemsbiology.cau.edu.cn/agriGOv2/) |
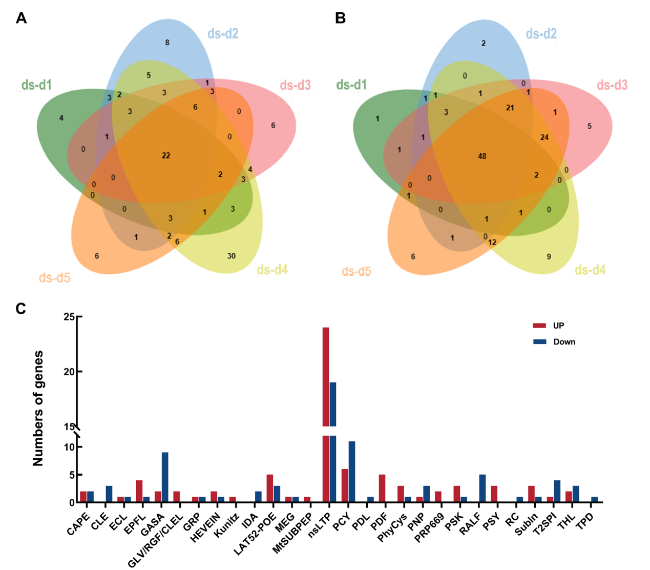
Fig. 5 Expression analysis of SlSSPs under drought stress. A Up-regulated SlSSPs in drought stress 1 day (ds-d1), 2 days (ds-d2), 3 days (ds-d3), 4 days (ds-d4) and 5 days (ds-d5) after treatments. Fold change = FPKMds-dn/FPKMck, Log2Foldchange ≥ 1 and qvalue ≤ 0.05. B Down-regulated SlSSPs in drought stress 1 day (ds-d1), 2 days (ds-d2), 3 days (ds-d3), 4 days (ds-d4) and 5 days (ds-d5) after treatments. Fold change = FPKMds-dn/FPKMck, Log2Foldchange ≤ -1 and qvalue ≤ 0.05. C Up- and down-regulated members of SlSSP families under drought stress |
Potential roles of tomato CEP family members in drought response
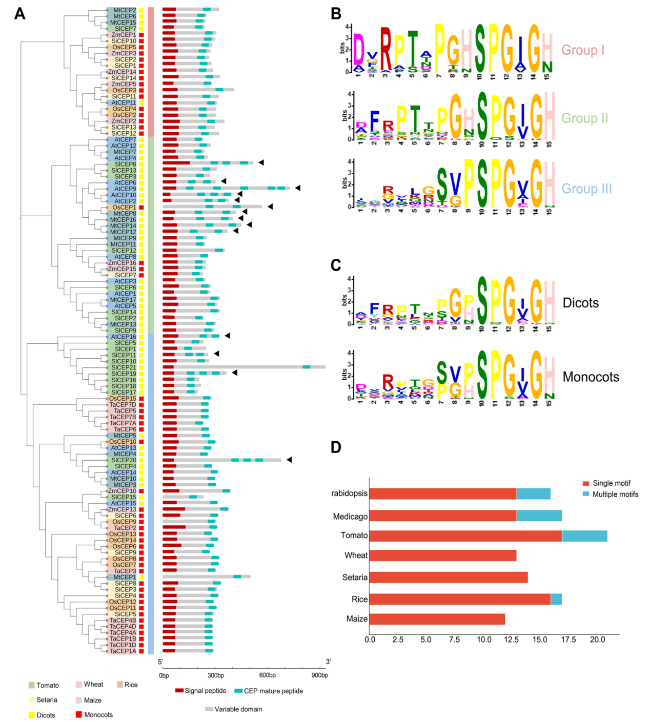
Fig. 6 Identification of CEP family members in tomato. A Phylogenetic tree and protein structure of AtCEPs, SlCEPs, MtCEPs, TaCEPs, OsCEPs, SiCEPs and ZmCEPs. The alignment was performed using Muscle with the full length protein sequences. The phylogenetic tree was constructed by the Maximum Likelihood method with 1000 bootstrap replications. The phylogenetic tree was drawn by the iTOL online tool (https://itol.embl.de/itol.cgi). The protein structures of CEPs were constructed by IBS software. Red boxes indicate monocotyledon plants, and yellow boxes indicate dicotyledon plants. Different colors indicate different species. Red boxes represent signal peptide, blue-green boxes represent CEP mature peptide, and gray boxes represent variable domain, “◀” represents CEP proteins containing multiple CEP domains in protein structure. B, C Logos were created by the MEME online tool (https://meme-suite.org/meme/tools/meme). D The number of single-domain genes (red) and multiple-domains genes (blue) in Arabidopsis, Medicago, wheat, Setaria, rice, maize and tomato |
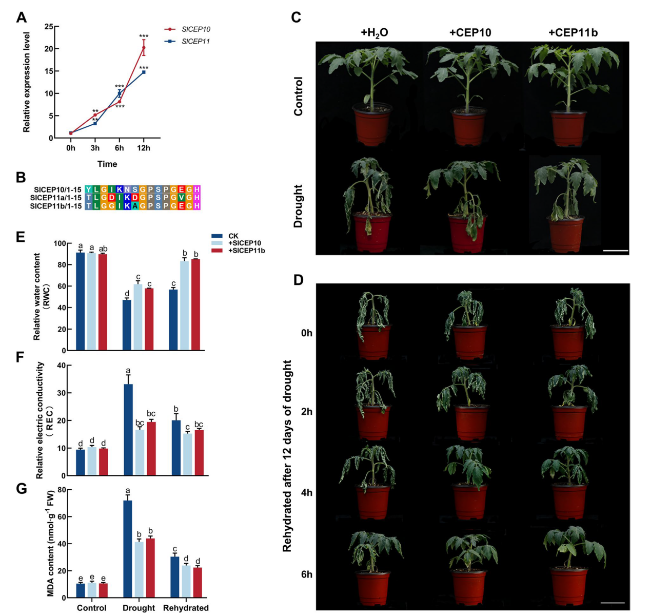
Fig. 7 Exogenous application of SlCEP10 and SlCEP11b peptides altered drought response in tomato. A Expression level of SlCEP10, SlCEP11 under drought (20% PEG6000) stress treatments for 0, 3, 6 and 12 h based on qRT-PCR. qRT-PCR was performed with three biological replicates and three technological replicates. The data represent mean ± SD, * p < 0.05, ** p < 0.01, *** p < 0.001 was determined by ordinary one-way ANOVA. * indicates a significant difference at the P < 0.05 level. B The sequences of synthesized SlCEP10, SlCEP11a and SlCEP11b peptides and their sequence alignment results. C 4-week-old tomato plants treated with 5 µM SlCEP10 or SlCEP11b peptide for 8 days under control and drought conditions. D 4-week-old tomato plants treated with 5 µM SlCEP10 or SlCEP11b peptide for 12 days under drought conditions and 1, 2, 4, 6 h after rehydration. E-G Measurement of relative water contents (RWC) (E), relative electric conductivity (REC) (F) and malondialdehyde (MDA) contents (G), respectively 8 days after drought condition. Sample size n = 15 in (E-G). All statistics analyses were performed with three biological replicates. Different lowercase letters in (E-G) indicate statistically significant differences based on ordinary one-way ANOVA (p < 0.05). Bar = 6 cm in (C, D) |
Exogenous application of SlCEP10 and SlCEP11b peptides enhanced drought tolerance of tomato plants
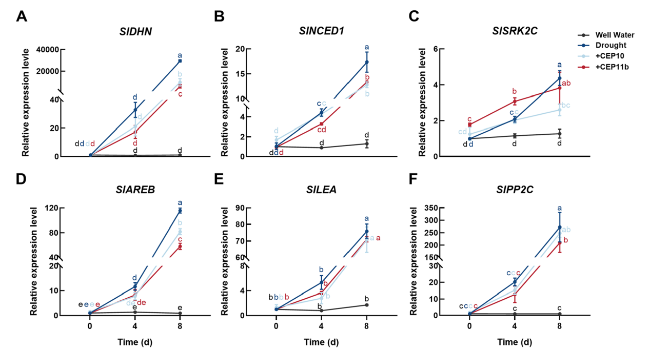
Fig. 8 Relative expression levels of drought-related genes under drought stress after exogenous application of SlCEP10 or SlCEP11b peptides. Relative expression levels of SlDHN (A), SlNCED1 (B), SlSRK2C (C), SlAREB (D), SlLEA (E) and SlPP2C (F) by qRT-PCR. qRT-PCR was performed with three biological replicates and three technological replicates. Different lowercase letters indicate statistically significant differences based on ordinary one-way ANOVA (p < 0.05) |