Introduction
APA dynamics and regulation
Dynamic features
Factors defining APA sites
APA dynamic in response to abiotic stress
Cd
Salt
Hypoxia and oxidative responses
Heat and cold
Circadian rhythms
APA dynamics in response to biotic stresses
Bacteria
Fungi
Viruses
Features of stress induced APA genes
Table 1 Dynamic features of plant APA genes related to abiotic stress responses |
| Abiotic stress | ||||||
|---|---|---|---|---|---|---|
| Stress | General trend of 3’UTR | Poly(A) site usage | Gene | Expression level | Reference | |
| Cd | / | proximal | AT5G54370(long-time stress); OsPrx1, OsRPA1a, OsEXO1 | ↑ | (Cao et al. 2019; Ye et al. 2019b; Zhou et al. 2019) | |
| AtBGLU23, AtTUB2, AtCAP1, AtPGY1, AtCSLE 1, AtRH19 | / | |||||
| distal | AtBT4, AtAPI2, AtHP60, AtCYP81F 4, AtARP1, AtNOG1- 1, AT5G54370 (short-time stress), AtLOS1, AtASK2, OsPOD, CPSF30 | / | ||||
| High salt | lengthening | distal | AT4G23050, AT5G62470, Thhalv10011676m, Thhalv10014879m | ↑ | (Ma et al. 2022; Wang et al. 2022) | |
| Hypoxia | / | non-3’ UTR | RBOHD, TPS8, GID1A, AT3G46530, AT3G23030, AT5G46780 | ↑ | (Cui et al. 2015; de Lorenzo et al. 2017; Ye et al. 2019b) | |
| distal | NADP-ME2, OsAPX1, OsAPX2 | ↑ | ||||
| DCA1, OsRZFP34 | ↓ | |||||
| Heat | lengthening | proximal | OsHSP74.8, OsHSP40, HSP101, | ↑ | (Wilkins et al. 2016; Ye et al. 2019b; Wu et al. 2020) | |
| distal | OsHSP82A, OsHsfB2c | ↑ | ||||
| OsCam1-1, LEA, OsPYL, RCAR3, OsSAPK5, OsJAZ8 | ↓ | |||||
| poly(A) tail dynamic: tail lengthen | HSP32, HSP21, HSP70, HSP70-3, HSP40, HSP101, HSP17.6II, HSP17.6C, AT1G59860, AT4G12400, AT3G09350, AT4G36040 | ↑ | ||||
| Cold | / | proximal | AT1G11280, AT3G13110 | ↑ | (Zhang et al. 2022) | |
| distal | AT5G53420 | ↑ | ||||
Table 2 Dynamic features of plant APA genes related to biotic stress responses |
| Biotic Stress | |||||||
|---|---|---|---|---|---|---|---|
| Species | Stress resistance related | Phenotype changing related | Reference | ||||
| Poly(A) site usage | Gene | Expression level | Poly(A) site usage | Gene | Expression level | ||
| Bacterium Xanthomonas oryzae pv. Oryzae | proximal | OsWAK14 | ↑ | distal | SGR, YL1, ASL2, OsFBK12 | ↓ | (Zhang et al. 2015; Ye et al. 2019b) |
| distal | OsPti1a, OsDR8, SRZ1 | ↓ | |||||
| OsSSI2 | / | ||||||
| Fungus Magnaporthe oryzae | proximal | OsJAZ1 | ↑ | / | (Shun et al. 2016; Ye et al. 2019b) | ||
| distal | OsAOC | ↓ | |||||
| Virus Rice strip virus | proximal | OsWRKY46, OsWRKY76, OsRAR1, OsRLCK107, OsDR8, DPF, POD (resistant rice) | ↑ | proximal | NYC1 | ↑ | (Sun and Jiang 2006; Ye et al. 2019b) |
| distal | OsNPR1, POD (susceptive rice) | ↓ | distal | YGL1 | ↓ | ||
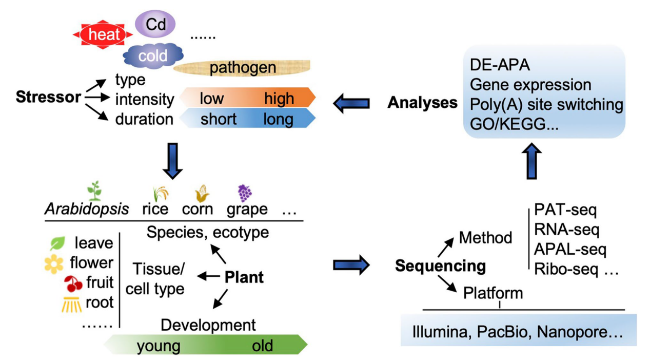
Fig. 1 Approaches to study stress induced APA dynamics. There are several points should be considered: stress type and intensity, plant tissue and developmental stage selections, and sequencing platform selection. The type of stress and its intensity can impact APA dynamics. The selection of plant tissue is crucial as APA dynamics often exhibit tissue and spatial heterogeneity. The choice of sequencing platform depends on the research objectives, and the analysis methods are based on the sequencing method |
Common mechanism of APA generation in response to stresses
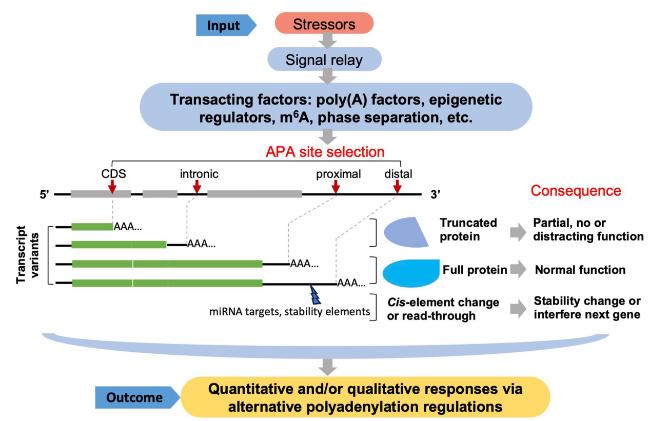
Fig. 2 A general model for transacting factors mediated stress responses through APA dynamics. Black lines represent gene sequences, thicker grey lines represent exons, red arrows point to poly(A) sites, “AAA…” represent the poly(A) tail. Poly(A) site switching to noncanonical regions, like at intronic or protein coding sequences (CDS) region, will alter the mRNA variants and produce a smaller truncated protein different from that of the full-length mRNA product. Proximal and distal poly(A) site switching within 3’UTR would shape the 3’ end profile without altering the protein sequences, but it is highly related to mRNA stability and/or gene expression level potentially through miRNA or stability factors. In some cases, transcription by-passing the normal poly(A) site may result in “read-through” into the next gene, which could cause gene silencing effect of the next gene. The impacts of APA on transcripts and gene expression are listed on the right, but not all consequences are applicable to all transcripts. The fate of individual kind of transcripts are circumstantially influenced by different factors |