During the editing of
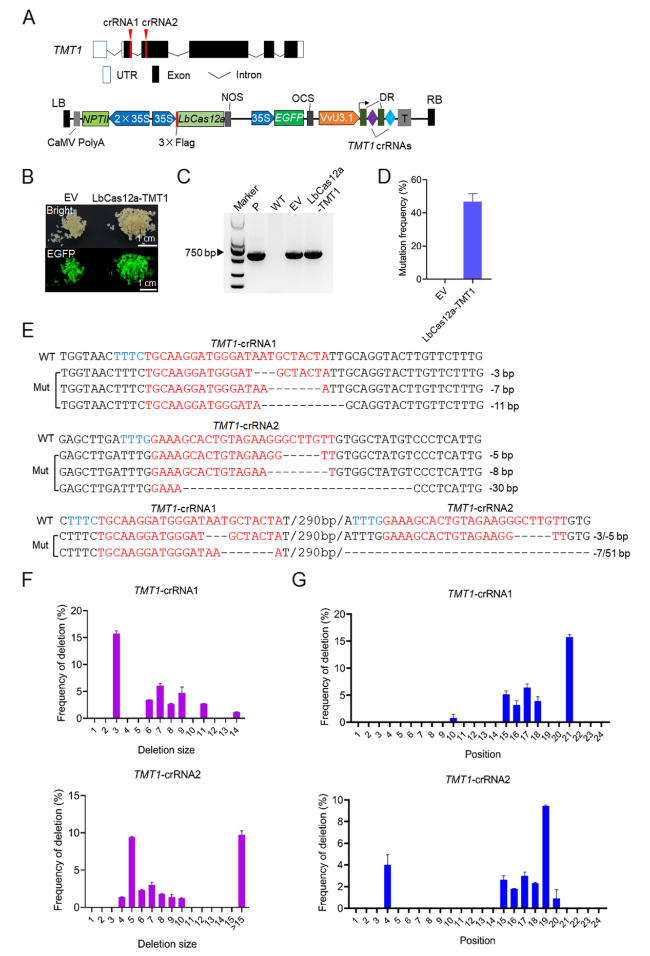
TMT1 gene, we found that the editing efficiencies for
TMT1-crRNA1 were always higher than that for
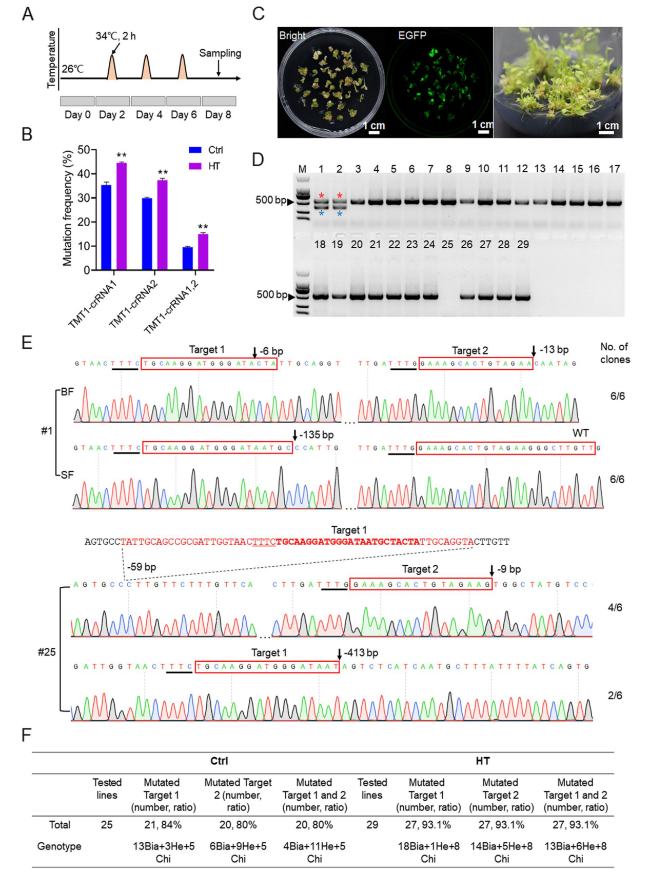
TMT1-crRNA2 before and after HT in grape cells (
Fig. 3B). Accordingly, the ratios of biallelic mutations for
TMT1-crRNA1 were also higher than that observed for
TMT1-crRNA2 in transgenic grapevine plants (
Fig. 3F). We therefore speculated that the editing efficiency might be affected by the difference of target sequences and/or the positions of crRNAs within the crRNA array. To test this hypothesis, we designed two crRNAs targeting the
TMT2 gene and combined them with previously developed
TMT1-crRNAs to make the
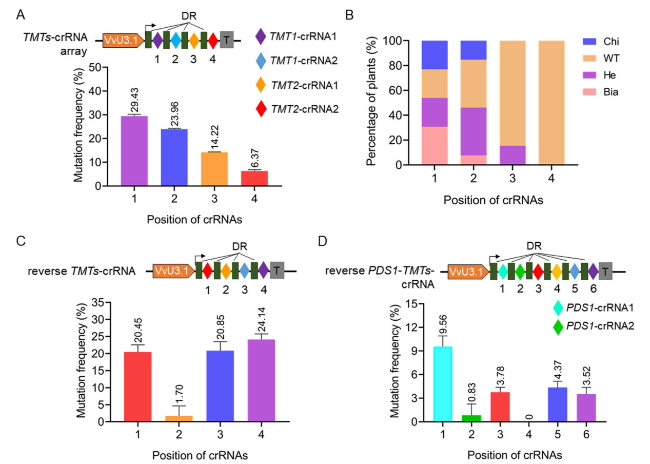
TMTs-crRNA array (
Fig. 4A, Supplementary Fig. S4). The resulting editing vector was introduced into 41B cells, and targeted mutagenesis was successfully detected at the four targets (Supplementary Fig. S4, Table S7). Simultaneous knockout of
TMT1 and
TMT2 resulted in reduced sugar accumulation in grape cells (Supplementary Fig. S4), which is consistent with the previous results obtained using CRISPR/Cas9 (Ren et al.
2021). The editing efficiencies with respect to the four individual targets were evaluated, and the results showed that the highest editing efficiency was achieved by using
TMT1-crRNA1 (29.43%), which is placed at position 1 (the most promoter-proximal crRNA) in the
TMTs-crRNA array (
Fig. 4A, Supplementary Table S7). On the contrary, the
TMT2-crRNA2 at position 4 resulted in the lowest editing efficiency (6.37%) (
Fig. 4A, Supplementary Table S7). Furthermore, analysis of 13 transgenic grapevine plants showed that the highest percentage of edited plants, including biallele mutants, heterozygotes, and chimeras, was observed with
TMT1-crRNA1, whereas no mutated plants were detected for
TMT2-crRNA2 (
Fig. 4B, Supplementary Fig. S5). To further confirm our hypothesis, we exchanged the order of crRNAs to make the reverse
TMTs-crRNA array without changing the target sequences (
Fig. 4C). If the editing is just influenced by the order of crRNAs, the editing profiles of the crRNAs at the same positions within the array are expected to be unchanged or less affected irrespective of the target sequences. However, the results obtained using the reverse
TMTs-crRNA array showed that the two crRNAs designed for
TMT1 still resulted in higher mutation frequencies (24.14% for
TMT1-crRNA1 at position 4 and 20.85% for
TMT1-crRNA2 at position 3) when compared with the two
TMT2-crRNAs (20.45% for
TMT2-crRNA2 at position 1 and 1.7% for
TMT2-crRNA1 at position 2) (
Fig. 4C, Supplementary Table S8). In addition, two
phytoene desaturase 1 (
PDS1) crRNAs were designed and added to the reverse
TMTs-crRNA array to develop the reverse
PDS1-
TMTs-crRNA array (
Fig. 4D, Supplementary Fig. S6). The newly developed crRNA array was introduced into 41B cells with LbCas12a and editing efficiencies were evaluated by using Hi-TOM assay. The editing profiles resulted by the four
TMTs crRNAs in the reverse
PDS1-
TMTs-crRNA array was similar to that using the reverse
TMTs-crRNA array as shown in
Fig. 4C, though the positions of the four crRNAs had been altered (
Fig. 4D, Supplementary Table S9). All these results indicated that the target sequence is the predominant factor affecting editing efficiencies during multiplex genome editing.