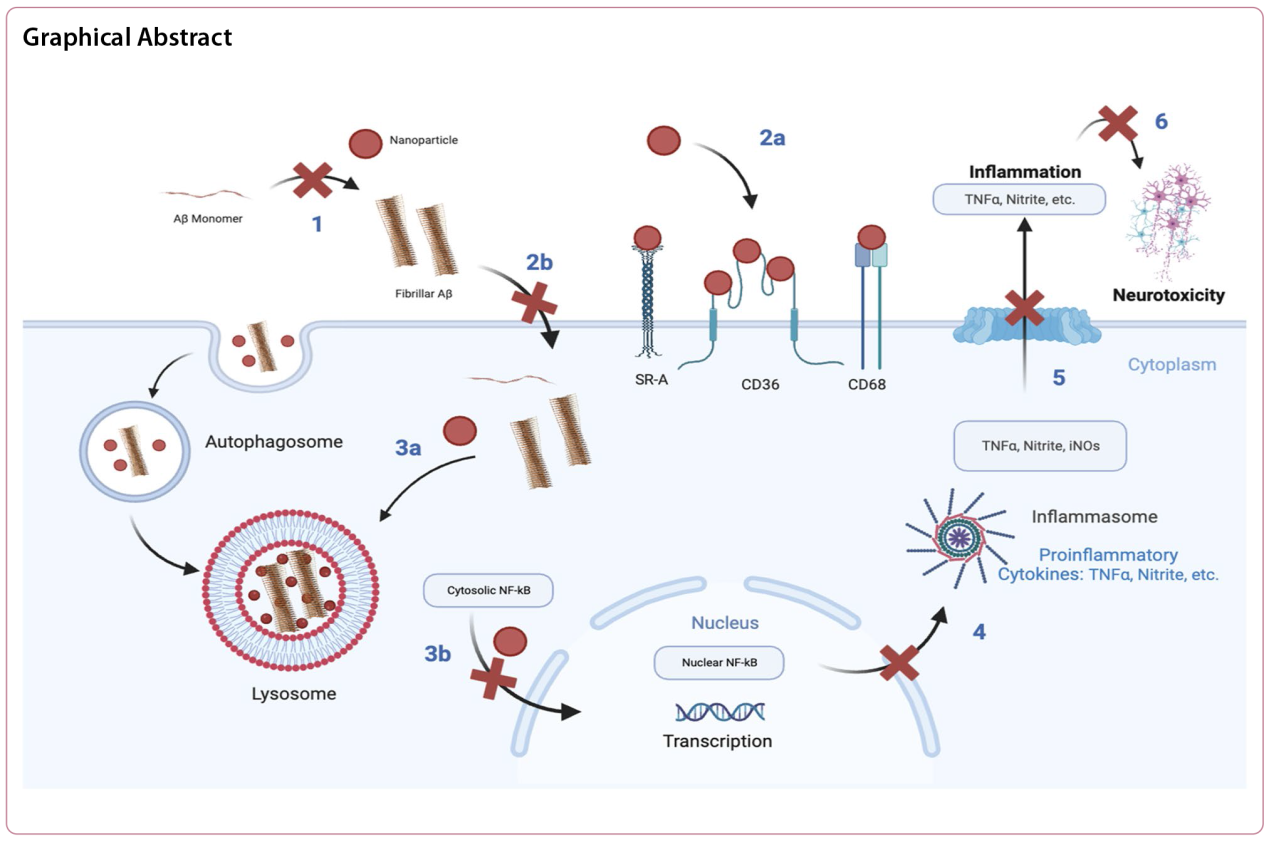
Background
Methods
Synthesis of AMs
Nanoparticle fabrication and characterization
Fig. 1 Nanoparticle fabrication via flash nanoprecipitation (FNP). a Schematic diagram of the FNP procedure for the synthesis of AM-nanoparticles (NPs). A confined impinging jet mixer (CIJ) was used to mix a stream of 250 µl of 50% (v/v) mixture of tetrahydrofuran (THF) containing 8 mg/ml shell molecule and 2.5 mg/ml hydrophobic core molecule with 250 µl of an aqueous stream. Key element of FNP technique for the fabrication of stable NPs is the induction time of aggregation and precipitation. The time required for a complete and homogenous mixing of the aqueous and solvent stream is Tmix, while the precipitation time of NPs is Tflash. b Table listing the chemical structures of AM shell molecules and polystyrene core. c Transmission electron microscope (TEM) images of NPs. d Table summarizing the characterization of NP radius and polydispersity index (PDI). Data are presented as mean ± SEM; n = 3 |
Preparation and characterization of Aβ1-42 fibrils
Th-T fluorescence assay
TEM
Kinetics of Aβ fibrillization
Cell culture
BV2 mouse microglia cell line
SH-SY5Y human neuroblastoma cell line
Cell-based competitive receptor binding assay
Aβ internalization assay
Thioflavin-S assay
Tumor necrosis factor-alpha (TNF-α) and nitric oxide (NO) assay
Immunocytochemistry
Lysosome activity
Autophagic activity
NF-κB translocation assay
Image acquisition
Neurotoxicity assay
Statistical analysis
Results
AM-NPs prevent Aβ fibrilization
Fig. 2 Development of fibril amyloid beta (fAβ) and the effect of NPs on Aβ fibrilization kinetics. a Representative transmission electron microscope (TEM) images of fAβ and oligomeric Aβ (oAβ). fAβ or oAβ (both 5 μl) was loaded on a formvar-coated, carbon-stabilized copper grid. Scale bar, 200 nm. b Thioflavin-T (Th-T) assay for fAβ and oAβ to validate the aggregation of β sheets. Protein or PBS (control) was mixed with 40 µM Th-T in a black-bottom 96-well microplate. Data are presented as mean ± SEM; n = 3; **P = 0.003 for student’s t test. c Th-T kinetic assay conducted with Aβ monomer over 65 h at 37 °C. Samples containing monomeric Aβ only or in the presence of NPs were loaded with 20 µM Th-T into a 96-well clear-bottomed non-binding half-area plate. d Endpoint Th-T fluorescence at 63 h. Data are presented as mean ± SEM; n = 3-4; ****P < 0.0001 for T12P5(PS) versus fAβ488 and ***P = 0.0007 for M12P5(PS) versus fAβ488 for Dunnett’s multiple comparisons shown on graph by one-way ANOVA. e TEM images of preformed fAβ mixed with AM-NPs or distilled water for 2 h. Scale bar, 500 nm |
AM-NPs bind to fAβ-binding SRs and interrupt fAβ internalization
Fig. 3 Competitive receptor binding assay in vitro. NPs reduce the surface active sites of CD36 (a), CD68 (b), SRA1 (c), and (d) TLR2 on the surface of BV2 microglia. BV2 microglia were incubated with NPs for 1 h, then co-incubated with SR-specific antibody or isotype controls for 30 min. After fixation, cells were washed, then blocked with 2% goat serum without PBS-T. To avoid internalization of NPs or antibodies, cells were incubated on ice until imaged. Cells were incubated with secondary antibodies including Alexa 488 or 594 for 1 h at room temperature. Data are presented as mean ± SEM; n = 3; ***P = 0.0001 for a, **P = 0.009 for b, **P = 0.0048 for c, Dunnett’s multiple comparisons shown on graph by one-way ANOVA |
Fig. 4 Scavenger receptors interrupt the microglial internalization of fibril amyloid beta (fAβ). BV2 microglia were treated with antibodies against scavenger receptors CD36, CD68, TLR2, and SRA1 or isotype control for 24 h, and then co-incubated with Alexa fluor 488-labeled fAβ for 24 h. The cells were fixed with 4% PFA, then washed two times with PBS to remove the extracellular fAβ. Cells were then incubated with 0.5% Triton-X-100 in potassium buffered saline (PBS-T) to remove any membrane-bound fAβ particles. a Representative confocal images of intracellular fAβ488 (green) and brightfield illuminated cells (grey). Scale bar, 50 µm. b Quantitative measurement of the intracellular Aβ488 in BV2 cells. Data are presented as mean ± SEM; n = 4; ****P < 0.0001 for Dunnett’s multiple comparisons shown on graph by one-way ANOVA |
Fig. 5 Effect of NPs on amyloid beta (Aβ) cellular uptake. BV2 microglia were treated with NPs for 24 h and then co-incubated with Alexa fluor 488-labeled fAβ for 24 h. Cells were fixed with 4% PFA and then washed two times with PBS to remove extracellular fAβ. Cells were then incubated with 0.5% Triton-X-100 in potassium buffered saline (PBS-T) to remove any membrane-bound fAβ particles. a Confocal images of intracellular Alexa fluor 488-labeled fAβ (green) and brightfield illuminated cells. Scale bar, 50 µm. b Quantitative measurement of the intracellular fluorescence of Aβ488 in BV2 microglia. Data are presented as mean ± SEM; n = 4; **P = 0.0016 for T12P5 (PS) versus fAβ488, and **P = 0.0059 for M12P5 (PS) versus fAβ488, for Dunnett’s multiple comparisons shown on graph by one-way ANOVA. c Thioflavin-S stain (blue) of fAβ in BV2 microglia (brightfield/grey) treated with NPs for 24 h, and then co-incubated with fAβ for 24 h. d Quantitative measurement of the intracellular fluorescence of Thioflavin-S stain in BV2 microglia. Data are presented as mean ± SEM; n = 4; ****P < 0.0001 for T12P5(PS) versus fAβ488, and ****P < 0.0001 for M12P5(PS) versus fAβ488, for Dunnett’s multiple comparisons shown on graph by one-way ANOVA |
AM-NPs modulate microglial inflammatory response and neurotoxicity
Fig. 6 AM-NPs suppress Aβ-mediated microglial activation. a-c BV2 microglia were treated with 20 µM Aβ in the presence or absence of NPs or 10 ng/ml lipopolysaccharide (LPS). After 24 h, supernatants were collected and assayed for inflammatory markers. Cells were fixed with 4% PFA and then permeabilized in 0.5% Triton-X-100 in potassium-buffered saline (PBS-T). After blocking, cells were incubated with anti-iNOS antibody overnight at 4 °C, then washed with PBS-T. Cells were incubated with secondary antibody Alex 594 for 1 h. a Confocal images representing the expression of inducible nitric oxide synthase (iNOS) in BV2 cells after 24-h co-incubation with fAβ and NPs. Scale bar, 25 µm. b Quantitative analysis of fluorescence intensity of iNOS in BV2 cells after 24-h co-incubation with fAβ and NPs. Data are presented as mean ± SEM; n = 3; ***P = 0.0002 for T12P5 versus fAβ, ***P = 0.0001 for M12P5 versus fAβ for Dunnett’s multiple comparisons shown on graph by one-way ANOVA. c Nitrite concentration measured by Griess reagent in the conditioned media harvested 24 h after cell co-incubation with Aβ and NPs. Data are presented as mean ± SEM; n = 3; **P = 0.0057 for T12P5 versus fAβ, **P = 0.0048 for M12P5 versus fAβ for Dunnett’s multiple comparisons shown on graph by one-way ANOVA. d Concentration of TNFα measured by ELISA in the conditioned media harvested 24 h after cell co-incubation with fAβ and NPs. Data are presented as mean ± SEM; n = 3; **P = 0.042 for T12P5 versus fAβ, **P = 0.0068 for M12P5 versus fAβ for Dunnett’s multiple comparisons shown on graph by one-way ANOVA. e SH-SY5Y cells were treated with conditioned media of fAβ- or LPS-stimulated BV2 cells in the presence or absence of NPs. Cytotoxicity in response to stimulated BV2 conditioned media was quantified in SH-SY5Y media using LDH assay. Data are presented as mean ± SEM; n = 3; ***P = 0.003 for T12P5 versus fAβ and **P = 0.0027 for M12P5 versus fAβ for Dunnett’s multiple comparisons shown on graph by one-way ANOVA |
AM-NPs modulate NF-kB nuclear translocation and induce lysosomal clearance
Fig. 7 NPs mediate fAβ-induced NF-ĸB nuclear translocation in microglia. a, b BV2 microglia were treated with 20 µM fAβ with or without NPs for 2 h. Cells were immediately fixed with 4% PFA. Cells were then blocked using 2% blocking buffer, incubated with anti-NF-ĸB P65 antibody overnight at 4 °C, and washed with PBS-T. Cells were incubated with secondary antibody Alex 488 for 1 h and nuclei were counterstained with Hoechst (blue). a Representative confocal microscopy images showing NF-ĸB nuclear translocation. Scale bar, 25 µm. b Quantitative measurement of the nuclear translocation of NF-ĸB in BV2 cells treated with Aβ with or without NPs for 2 h. Data are presented as mean ± SEM; n = 3; **P = 0.0027 for fAβ versus control, **P = 0.0044 for T12P5 versus fAβ and **P = 0.0054 for M12P5 versus fAβ, for Dunnett’s multiple comparisons shown on graph by one-way ANOVA |
Fig. 8 AM-NPs accelerate lysosomal clearance of fAβ. a-d BV2 microglia were incubated with NPs for 24 h and then co-incubated with Alexa Fluor 488-labeled fAβ for 2 h or 24 h. Lysosomes in treated live BV2 microglia were stained with 70 µM Lysotracker (red) for 30 min. Cells were then fixed using 4% PFA. a Representative confocal microscopy images showing fAβ-positive (green) lysosomes (red). White arrows show fAβ in lysosomes. Scale bar, 25 µm. b Quantitative measurement of the lysosomal degradation of intracellular Aβ at 2 h and 24 h in BV2 microglia pretreated with NPs. Data are presented as mean ± SEM; n = 3; ****P < 0.0001 for T12P5 versus fAβ, for Dunnett’s multiple comparisons shown on graph by two-way ANOVA. c, d BV2 microglia incubated with NPs for 24 h then co-incubated with Alexa Fluor 488-labeled fAβ for 2 h. Cells were fixed with 4% PFA, permeabilized using PBS-T, and then blocked using 2% goat blocking buffer. Cells were then incubated with anti-CD68 or LAMP-1 antibody overnight at 4 °C, then washed with PBS-T. Cells were incubated with secondary antibody Alex 488 or 954 for 1 h and nuclei were counterstained with Hoechst. c Orthogonal projection of confocal Z-stacks shows labeling of CD68 (green), Dil-labeled NPs (red), and nuclear stain Hoechst (blue). d Orthogonal projection of confocal Z-stacks shows labeling of LAMP-1 (white), Dil-labeled NPs (red), nuclear stain Hoechst (blue), and fAβ488 (green) in brightfield illuminated cells in the top panel, and co-labeling of nuclear stain Hoechst (blue), Dil-labeled NPs (red), and fAβ488 (green) in brightfield illuminated cells in the bottom panel. White arrows show intracellular fAβ. e, f BV2 microglia were incubated with NPs for 24 h and then co-incubated with fAβ for 30 min. e Confocal images of LC3 immunoreactivity (green) in BV2 microglia co-stained with Hoechst (blue). f Quantitative analysis of LC3 fluorescence intensity in BV2 microglia. Data are presented as mean ± SEM; n = 3; **P < 0.01 for T12P5 versus fAβ, for Dunnett’s multiple comparisons shown on graph by two-way ANOVA. Scale bar, 25 µm |
Fig. 9 AM-NPs accelerate clearance of fAβ. a-d Electron microscope micrographs of BV2 microglia incubated with NPs for 24 h, then co-incubated with fAβ for 30 min. Cells were fixed with a mixture of 2.5% glutaraldehyde and 4% PFA in 0.1 M cacodylate buffer at pH 7.4. Ultrathin (90 nm) sections of cell pellets were imaged using the electron microscope. a, e The ultrastructure of a microglia treated with fAβ only. The intracellular fAβ (white arrowhead) is surrounded by damaged vacuoles, including large autophagosomes (white arrows) and lysosomes with incomplete membranes (yellow arrows). Mitochondria (M) in this cell is structurally damaged, as evidenced by disordered cristae and a thin outer membrane (e). b, f The ultrastructure of a fAβ-treated microglia co-incubated with T12P5(PS). In this cell, the phagocytic activity is high, as illustrated by the high representation of a variety of autophagosomes (white arrows), lysosomes with well-developed membranes (yellow arrows), and autophagosome-lysosome fusion structures (blue arrows). Mitochondria in this cell are intact, as evidenced by well-defined cristae structures and an intact outer membrane (f). c, g The ultrastructure of fAβ-treated microglia co-incubated with M12P5(PS). The phagocytic activity is less, as illustrated by fewer autophagosomes (white arrows); however, the lysosomal structures are characterized by a well-developed double membrane (yellow arrows). This cell has a high representation of intact and long mitochondria characterized by well-defined cristae and an intact outer membrane (g). d, h The ultrastructure of fAβ-treated microglia co-incubated with the control PS-b-PEG(PS). A large center of fAβ is present in this cell, along with large thin-walled autophagosomes (white arrows) and thin-walled lysosomal structures (yellow arrows). Mitochondria in this cell are structurally damaged, as evidenced by distorted cristae and thin outer membrane. N, nucleus, and M, mitochondria. Scale bars, 500 nm in the top panels of a-d and e-h; 1 µm in the bottom panels in a-d |