Introduction
Results
Fluorescence imaging deformability cytometry (FI-DC) platform
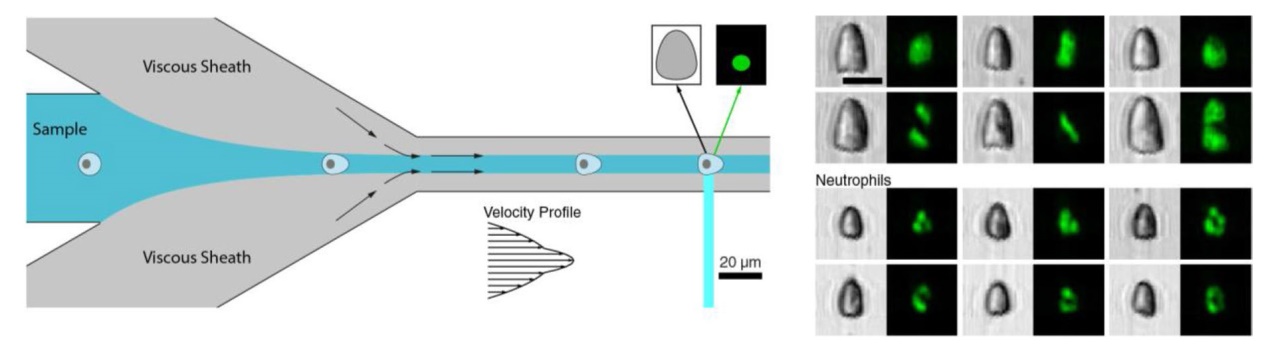
Fig. 1 FI-DC schematic and analysis overview. a Cells flow through a microfluidic device with viscous sheath flows that create a pinched velocity profile (inset). After deforming, cells are interrogated in the straight channel via FIRE, producing brightfield and fluorescence images. b Jurkat and neutrophil images demonstrating varied deformed shapes and nuclear structure. c Cells and nuclei are segmented from images using U-Net neural networks. Curvature is measured from extracted boundaries, which is integrated to yield a scale-invariant bending factor, which we use for cell deformability. Chromatin content is calculated as the ratio of nuclear and cell area. d Cell area, cell deformability, and chromatin content capture fundamental features of Jurkat cell populations (n = 10,031). Unlabeled scale bars are 10 μm |
Deformation characterization
Fig. 2 FI-DC features tunable deformation and reveals nuclear and membrane details. a Increasing the sample stream viscosity and the ratio of sample:sheath viscosity increases visible deformation. b Bending factor as a form of cell deformability captures the change in shape visible in images (n ≥ 11,700 for all groups). c Nuclear images of Jurkat cells reveal a subpopulation with reduced chromatin content (n = 9,730). d This reduced chromatin content subpopulation features a higher cell size and deformability than the rest of the cell population. e When comparing cells of similar size (blue box with 5 μm2 width in Fig. 2d), there is not a noticeable difference in cell deformability (n = 1,783). f Jurkats of similar size with low (yellow) and high (blue) cell deformability are selected for comparison (n = 10,031). g These cells do not exhibit noticeably different levels of chromatin content (blue n = 96; yellow n = 99). h Median shapes of cells in these populations reveal the high cell deformability cells are deformed more in the lower (back) corners of the cell. i Differences in the deformation of sections of the cell quantitatively demonstrate the sections of the cell responsible for overall cell deformability differences. Unlabeled scale bars are 10 μm |
Revealed nuclear structure heterogeneity
Investigating differences in cell deformability
Neutrophil extracellular traps
Fig. 3 FI-DC reveals changes in NETosing neutrophil deformation and structure. a Stimulated neutrophils from two donors have increased cell deformability and reduced chromatin content. Traces from 5 cells from each condition reveal cells have increased in size (n ≥ 21,700 for all groups). b The nuclear bending factor of stimulated neutrophils is slightly lower, as the cells begin the NETosis process. c There is no visible correlation between cell deformability and the shape of nuclear structure (nuclear bending factor) |
Fig. 4 NETosing neutrophils have increased cell deformability. a NETosing neutrophils demonstrate a noticeable increase in cell size and deformability (n ≥ 21,700 for all groups). b The cell deformability of neutrophils from a narrow size range is compared, which illustrates a small shift in deformability over this small size range (n ≥ 4,600 for all groups). c Distributions of cell deformability are normalized, which also show an increase in treated neutrophils. Bayesian analysis indicates that there is a significant change in the distributions’ means when modeled as t-distributions |