Introduction
Toehold-mediated strand displacement reactions
Working principle of TMSDRs
Fig. 1 Schematic representation of toehold-mediated strand displacement reaction. DNA is drawn as directional lines where the sequence information is omitted, and the arrow indicates 5' to 3' direction of the backbone. The sequence is divided into different color domains that have specific functions, while * indicates a complementary domain. a A forward reaction takes place only in the forward direction such reactants get converted into products. The ssDNA oligonucleotide U initiates the displacement of the strand S from the ST complex (also known as a duplex) by binding with the complementary toehold domain (α*). In the subsequent steps, the strand S is displaced through branch migration, forming |
Design considerations for TMSDRs
Computational and mathematical tools for analyzing TMSDRs
TMSDRs with different input and output signals
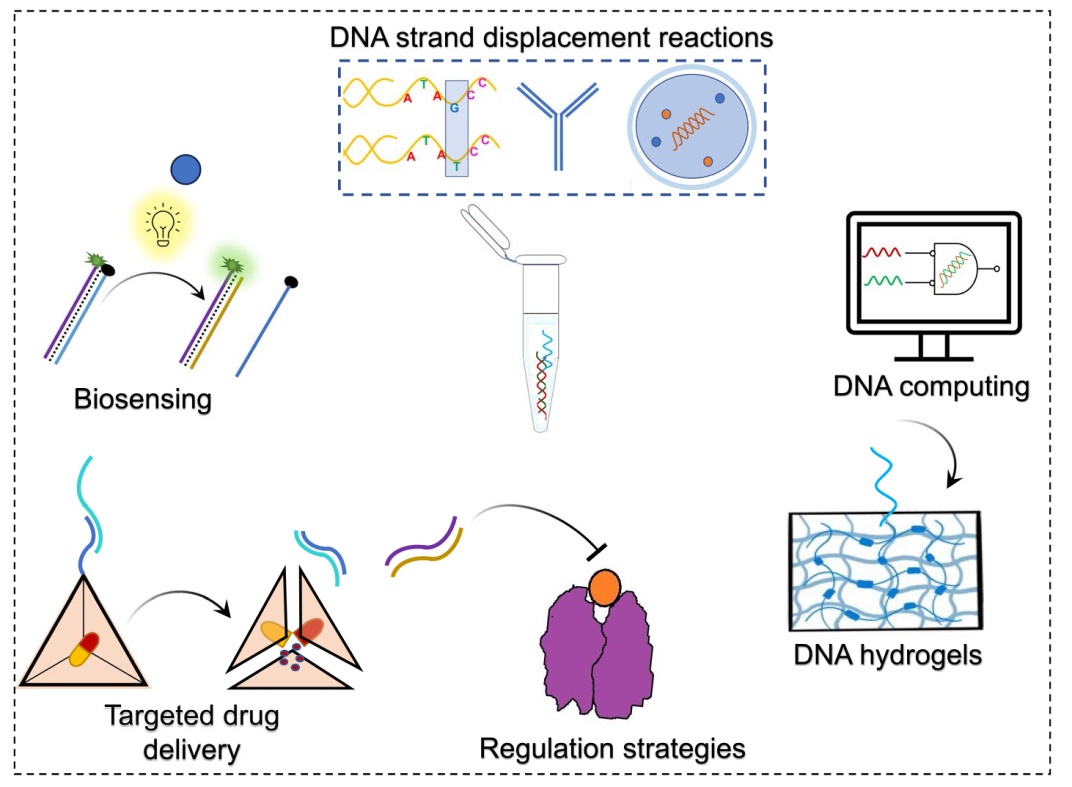
Fig. 2 a TMSDRs have been used to develop highly sensitive biosensors to detect clinically relevant targets, programmable drug delivery carriers for targeted therapeutics, regulation strategies to control the activity of clinically relevant molecules, responsive soft biomaterials like smart DNA hydrogels and DNA computing based applications. b Processing different input and output signals through TMSDRs [31]. Various chemical and physical signals such as adenosine triphosphate (ATP), pH, light, temperature, and electric signals can be converted to ssDNA while this information can be processed and produced in the form of diverse output signals such as change in fluorescence using fluorescence quenching mechanism and fluorescence resonance energy transfer (FRET), colorimetric method, gel-electrophoresis and atomic force microscopy (AFM) |
Processing chemical and physical input signals
Fig. 3 Examples of diverse input and output signals used by TMSDRs. a ATP as a signaling input. Here, a TMSDRs is initiated by ATP to produce a fluorescence signal through a ssDNA output. Reprinted with permission from Ref. [36], used under Creative Commons CC BY 4.0 license. b Temperature as a signaling input. Here, an increase in temperature above the melting point (Tm) opens the hairpin loop structure and releases a ssDNA output that can be used downstream in further cascaded reactions. Reprinted with permission from Ref. [37], used under Creative Commons CC BY license. c Colorimetric based output signal. Here, only in the presence of an input signal, a reporter-output complex can form that allows gold nanoparticles (AuNPs) to agglomerate, resulting in a detectable color change compared to dispersed AuNPs. Reprinted with permission from Ref. [38], used under Creative Commons CC BY 3.0 license. d Gel-electrophoresis based output signal. Here, the binding of HIV gene with a substrate probe (SP) results in an SP + HIV complex that has a higher molecular weight than SP alone. With the help of a fuel strand (FS), signal amplification can lead to higher fluorescence intensity. Reprinted with permission from Ref. [39], Copyright 2019, Royal Society of chemistry |
Producing diverse output signals
Biomedical applications of toehold-mediated strand displacement reactions
Applications of TMSDRs in biosensing
Detection of single nucleotide polymorphisms and variants
Fig. 4 Applications of TMSDRs in biosensing. a Use of TMSDRs for SNVs detection. Here, an X-shaped DNA probe with a quencher and fluorophore molecules disassociates when the complementary target strand binds with the DNA probe [63]. Further addition of two ssDNA probes, AP1 and AP2, releases the fluorescence tagged strand and the target strand to undergo multiple cycles of TMSDRs to amplify the fluorescence signal. Reprinted with permission from Ref. [63], copyright 2017, Elsevier B.V. b Use of TMSDRs for miRNA detection. Here, the target miRNA binds to the toehold and causes the displacement of the DNA walker probes, which further hybridize with a ferrocene-tagged signal probes [65]. The latter is released from the system using endonuclease, resulting in a detectable fluctuation in the oxidation current of the ferrocene. Reprinted with permission from Ref. [65], copyright 2020, American Chemical Society. c Use of TMSDRs for whole cell detection. Here, a hydrogel is functionalized with a DNA aptamer that can attach to specific cells during a dynamic flow [66]. The attached cells are programmed to be released at certain physiological conditions using strand displacement for further assay. Reprinted with permission from Ref. [66], copyright 2015, American Chemical Society. d Use of TMSDRs for real-time in situ detection. Here, a modular exchange network can detect dynamic changes in the target protein using a cascaded TMSDRs [23]. Reprinted with permission from Ref. [23], copyright 2020, Oxford University Press. e Use of TMSDRs driven DNA hydrogel for biosensing. Here, DNA oligonucleotide with aptamer-toehold sequence assists in specific binding to the cancer cells [67]. Two hairpin loops bind to the toehold and undergo a chain reaction to form hydrogel around the cell. Reprinted with permission from Ref. [67], copyright 2017, American Chemical Society |
Detection of MicroRNAs
Detection of whole cell
Real-time in situ detection mechanisms
Toehold-triggered DNA hydrogels for biosensing
Employing other techniques to enhance sensitivity and stability of TMSDRs based biosensors
Applications of TMSDRs in therapeutics
Targeted drug delivery
Fig. 5 Applications of TMSDRs in targeted drug delivery and regulation therapies. a Use of TMSDRs driven DNA nanomachine for transmembrane drug delivery. Here, DNA strands on the nanoparticles bind to input 1, a transmembrane receptor, which is over-expressed in the cancer cells, leading to endocytosis of the DNA nanomachine [115]. Hybridization of the toehold with intracellular miRNA, which is input 2, leads to the activation of phototherapy inside the cell. Reprinted with permission from Ref. [115], copyright 2021, American Chemical Society. b Use of TMSDRs driven DNA nanostructures for targeted drug delivery. Here, interaction with the target strand disassembles the DNA tetrahedron, which can be used for controlled drug release [117]. Reprinted with permission from Ref. [117], used under Creative Commons CC BY-NC 3.0 license. c Use of TMSDRs for regulating gene transcription. Here, complementary ssDNA and ssRNA reassociate on mutual interaction with another hybrid and release NF-kB decoys that block NF-kB proteins, thereby inhibiting gene expression [54]. Reprinted with permission from Ref. [54], used under Crown copyright. d Use of TMSDRs for regulating protein activity. Here, a regulatory circuit inhibits excess thrombin using a threshold controller network, which is designed to release inhibitor strands through TMSDRs that suppress thrombin activity when it is above a threshold [118]. Reprinted with permission from Ref. [118], copyright 2012, American Chemical Society. e Use of TMSDRs driven DNA hydrogel for targeted drug release. Here, AuNPs are functionalized with thiol groups to bind with DNA strands, which is designed to hybridize with other initiator strands and polymerize to form a DNA network hydrogel [119]. The target ligand binds to the toehold switch and disassembles the DNA shell in a controlled manner to release the drug. Reprinted with permission from Ref. [119], copyright 2019, American Chemical Society |
Regulating clinically relevant molecules
Toehold-triggered DNA hydrogels for therapeutics
TMSDRs driven DNA computing for biomedical applications
Fig. 6 Applications of TMSDRs based DNA computing. a DNA computing for biosensing. Here, a multi-input cascaded network can detect 4 serotypes of dengue. When Dengue A and DEN- k (k = 1,2,3,4) are present, the chimeric substrate is cleaved by the DNAzyme, producing an activator strand for cascaded signaling. Reprinted with permission from Ref [127], copyright 2014, Wiley-VCH. b DNA computing for targeted drug delivery. Here, multiple cancer cell surface markers are tagged with DNA aptamers, enabling TMSDRs to determine cellular viability and build a targeted photodynamic therapeutic device. The AND operation is activated by the first input aptamer, which displaces the incumbent strand and begins the cascaded layer reaction by exposing the second toehold. Reprinted with permission from Ref [132], copyright 2015, American Chemical Society. c-d DNA computing for programmable networks; c Here, the digital display works inside a well-plate and the logical circuit uses DNAzyme to function. The Boolean gate is operated in such a way that in the presence of an input strand, the FRET is cleaved by DNAzyme, releasing the FRET labelled DNA strand. Reprinted with permission from Ref [133], copyright 2014, Wiley-VCH; d Here, the fuel strand hybridizes with the target strand, and the resulting complex forms a waste product of lower energy. The consumption of the fuel strand causes the release of the target strand, which spontaneously hybridizes with the high-energy output strand. Reprinted with permission from Ref. [134] under creative common By-NC 4.0 license |