Introduction
Materials and methods
Datasets
Table 1 The detailed composition of three siRNA datasets. It shows the number of siRNAs and target mRNAs in each dataset, as well as corresponding publishers. Two siRNAs are removed in DT due to the limitation of i-score website, and two are removed in DR as a result of lacking binding sites on reported mRNAs |
 |
The architecture of inhibition predictor
Fig. 1 The architecture of our self-attention-based siRNA inhibition predictor. It is composed of three modules: feature extraction module, selfattention module, and fully connected module |
Feature extraction module
Self-attention module
Fully connected module
The architecture of our approach
The mRNA searching package
The off-target filter
The flow of our approach
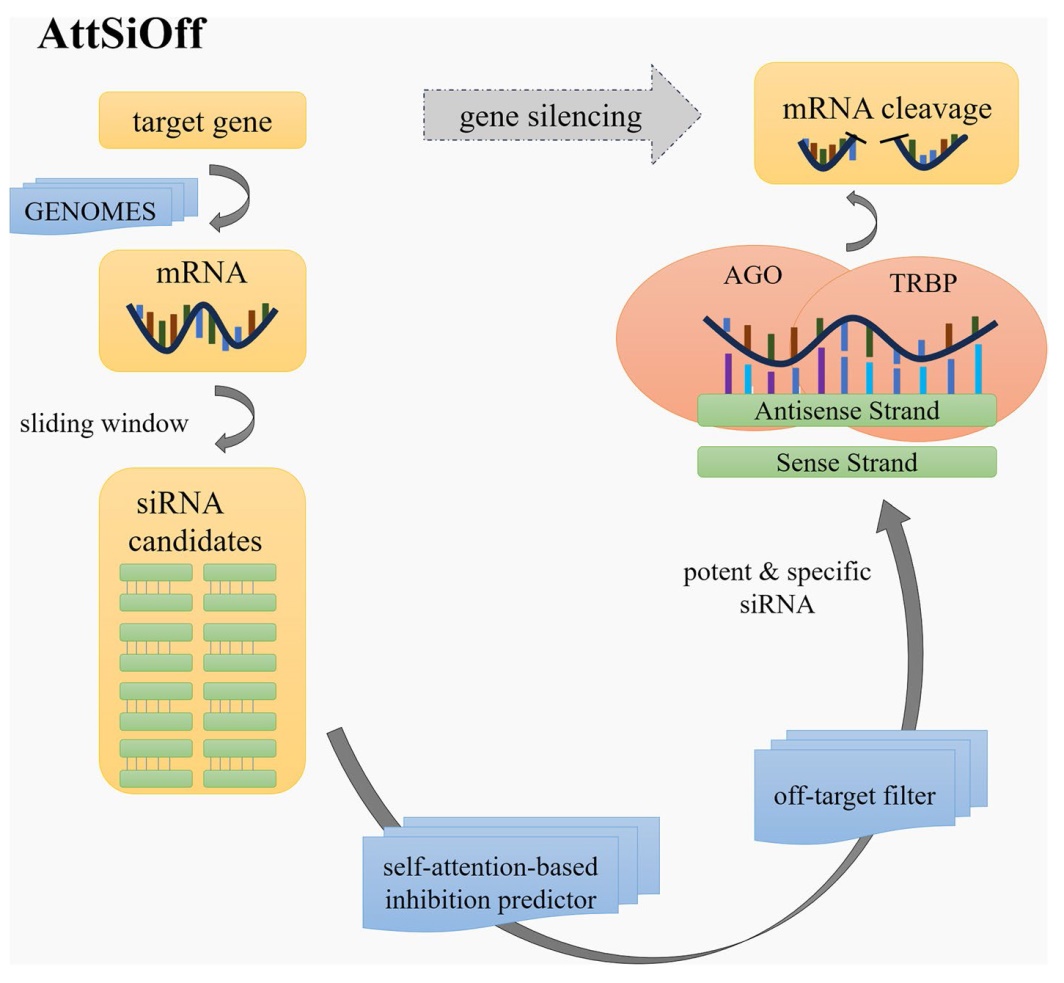
Fig. 2 The architecture of our siRNA design approach. It contains optional mRNA searching package pyGB, our siRNA inhibition prediction model, and optional off-target filter. The package pyGB can search mRNA sequences for given gene name from Genomes database. But one can also provide mRNA sequences directly. Substring and imprvoed Smith-Waterman algorithm are used to analyze on-target or miRNA-like offtarget effect, by aligning siRNA with each untargeted mRNA from Refseq or TargetScan 3'UTRs database |
Experimental setup
Evaluation metrics
Results and discussion
Tenfold cross-validation result
Fig. 3 The tenfold cross-validation result of our predictor compared with existing methods. Three metrics are shown with boxplots, including (a) PCC, (b) SPCC, and (c) AUC. The boxplot is used to describe 4-quartiles of data distribution. The orange lines represent mean values, the green dashed lines represent median values, and small circles represent outliers. Obviously, our method outperforms all existing methods among three metrics, reaching an average PCC of 0.81, SPCC of 0.84, and AUC of 0.886 |
Cross-dataset validation
Table 2 The cross-dataset prediction results. Bold numbers indicate the best results. Our method outperforms others on DT, while shows slight advantage on DR |
 |