

Journal of Diagnostics Concepts & Practice ›› 2023, Vol. 22 ›› Issue (05): 429-440.doi: 10.16150/j.1671-2870.2023.05.003
• Original articles • Previous Articles Next Articles
QIN Xiaodan, SUN Huiling, PAN Bei, PAN Yuqin, WANG Shukui( )
)
Received:2023-10-23
Online:2023-10-25
Published:2024-03-15
CLC Number:
QIN Xiaodan, SUN Huiling, PAN Bei, PAN Yuqin, WANG Shukui. miR-1229-3p inhibits the malignant progression of colorectal cancer and serves as a potential biomarker[J]. Journal of Diagnostics Concepts & Practice, 2023, 22(05): 429-440.
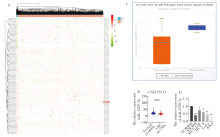
Figure 1
Downregulation of miR-1229-3p expression in human CRC cells A: Stratified clustering heatmap of differentially expressed miRNAs obtained from RNA sequencing data from CRCs and their corresponding normal tissues in the TCGA database. Downregulated genes were represented in red, upregulated genes in green and the red line represented miR-1229-3p; B: Expression of miR-1229-3p in GSE115513 dataset (normal tissue=649, tumor tissue=750); C: The expression of miR-1229-3p in the tissues of 450 colon cancer patients and 8 normal tissues in the Starbase database; D: qRT-PCR was used to detect the expression level of miR-1229-3p in human colorectal cancer cell line and human normal intestinal epithelial cell line NCM460. *:P<0.05; **:P<0.01; ***:P<0.001.
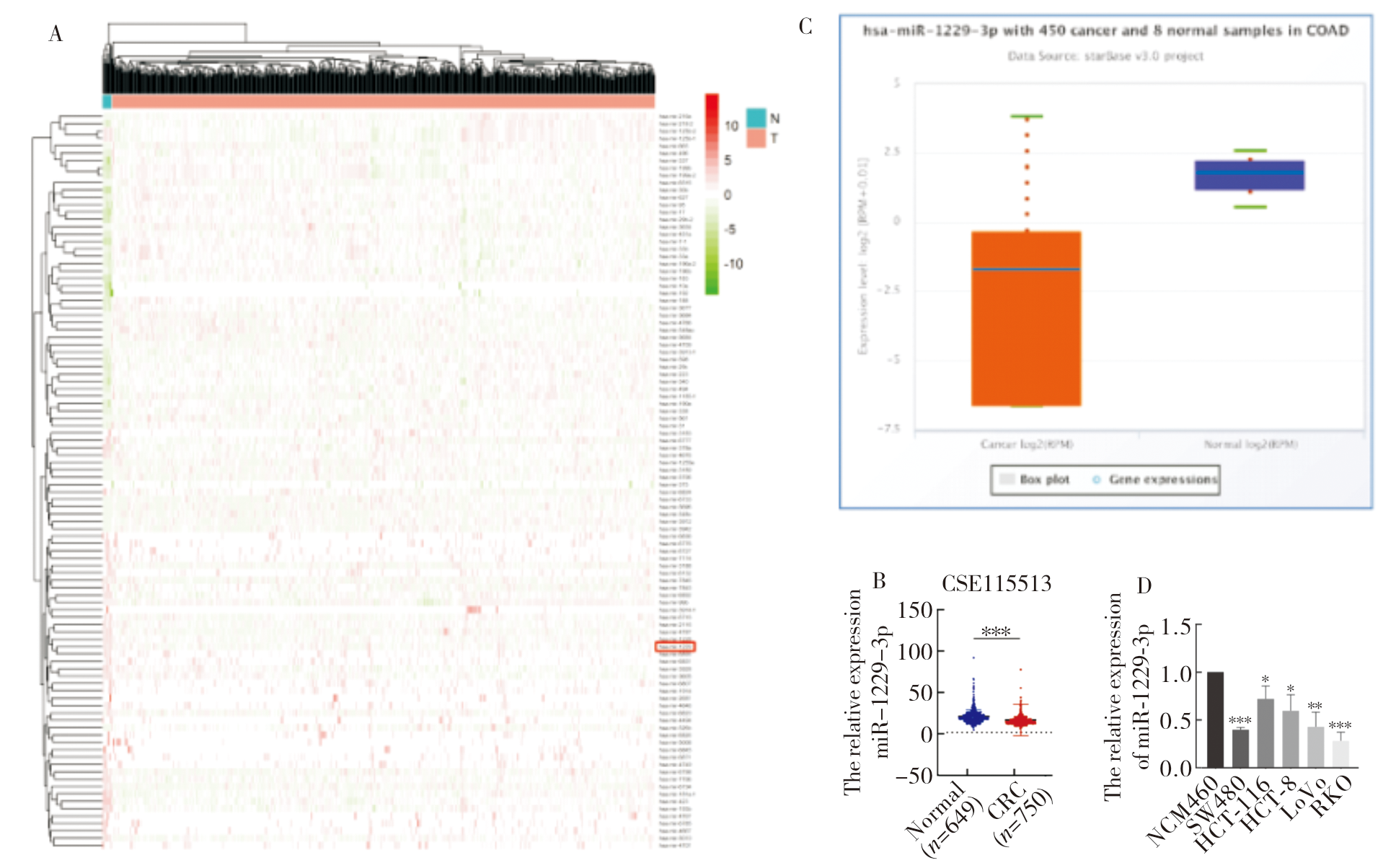
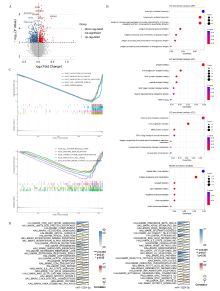
Figure 2
Bioinformatics analysis of miR-1229-3p biological functional characteristics A: Volcano plot showed the differential genes between the high miR-1229-3p expression group and the low miR-1229-3p expression group; B: GO and KEGG analysis of 725 differential genes; C: GSEA of miR-1229-3p based on GO and KEGG gene sets; D: Correlation analysis of miR-1229-3p with 50 marker pathways by GSVA. *:P<0.05; **:P<0.01; ***:P<0.001.
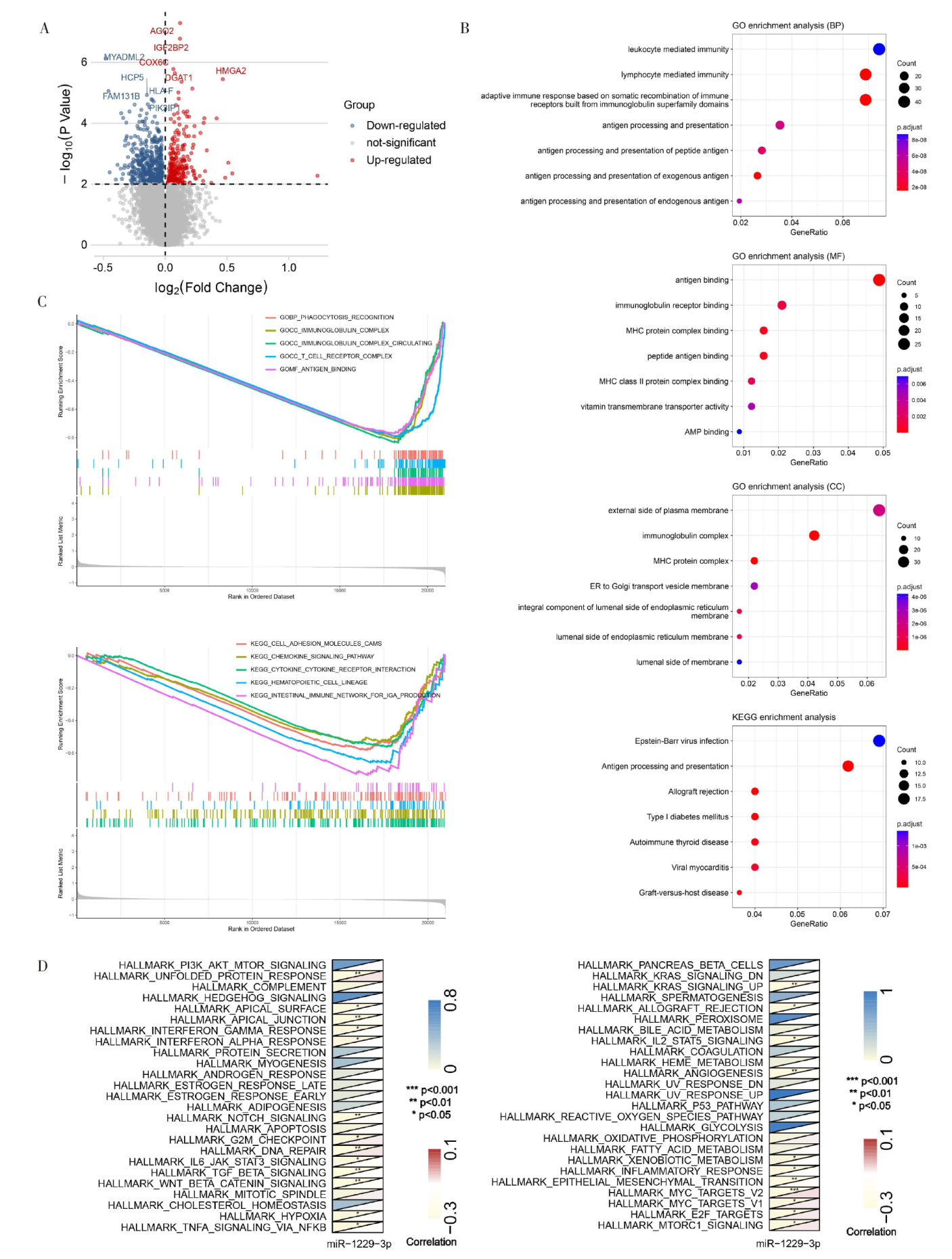
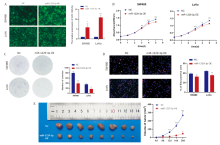
Figure 3
miR-1229-3p affected CRC cell growth A: Stable transfection SW480 and LoVo cells overexpressed by miR-1229-3p were constructed; B: CCK8 assay was used to explore the effect of miR-1229-3p overexpression on the growth ability of CRC cells; C: Clone formation assay was used to explore the effect of the overexpression of miR-1229-3p on the cloning ability of CRC cells; D: The effect of miR-1229-3p overexpression on the proliferation of CRC cells was explored by EdU assay; E: Tumor growth volume curves of tumor-bearing mice in the NC group and miR-1229-3p OE group. *:P<0.05; **:P<0.01; ***:P<0.001.
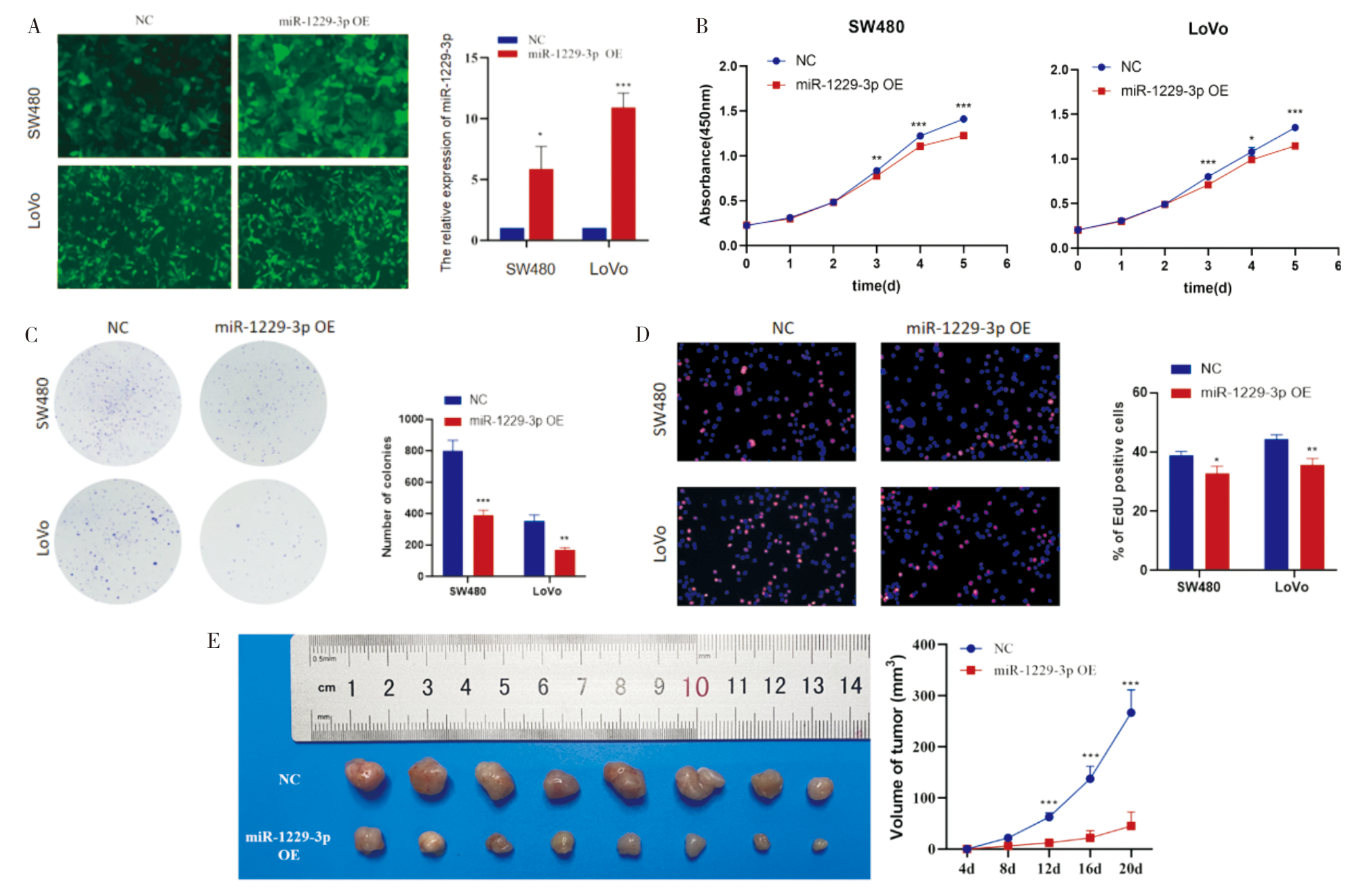
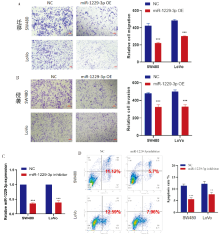
Figure 4
miR-1229-3p affected CRC cell migration, invasion, and apoptosis A: Transwell assay was used to explore the effect of miR-1229-3p overexpression on the migration of CRC cells; B: Transwell assay was used to explore the effect of miR-1229-3p overexpression on the invasion ability of CRC cells; C: Construction of miR-1229-3p-knockdown SW480 and LoVo cells; D: Flow cytometry assay was used to explore the effect of miR-1229-3p-knockdown on the apoptosis of CRC cells. **:P<0.01; ***:P<0.001.
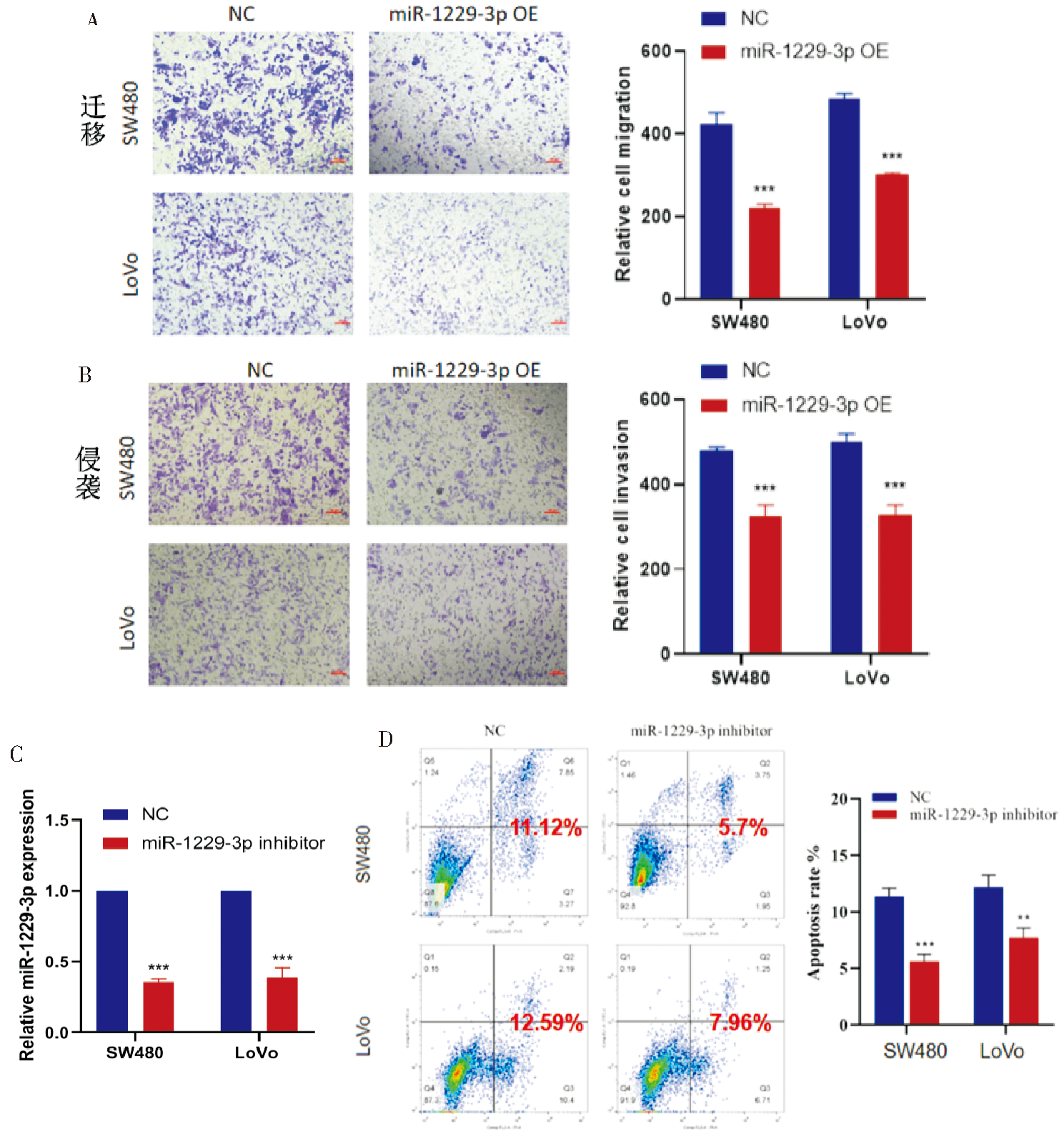
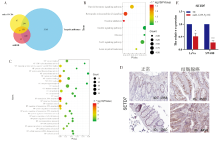
Figure 4
SETD7 was a potential target gene for miR-1229-3p A: The target gene of miR-1229-3p was predicted by DIANA, TargetScan, and miRDB databases; B: KEGG enrichment analysis of miR-1229-3p target gene; C: GO analysis of miR-1229-3p target gene; D: Expression of SETD7 in CRC from the Human Protein Atlas database; E:qRT-PCR was used to measure SETD7 in SW480 and LoVo cells when miR-1229-3p was overexpressed. *:P<0.05; ***:P<0.001.
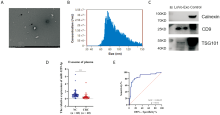


Figure 6
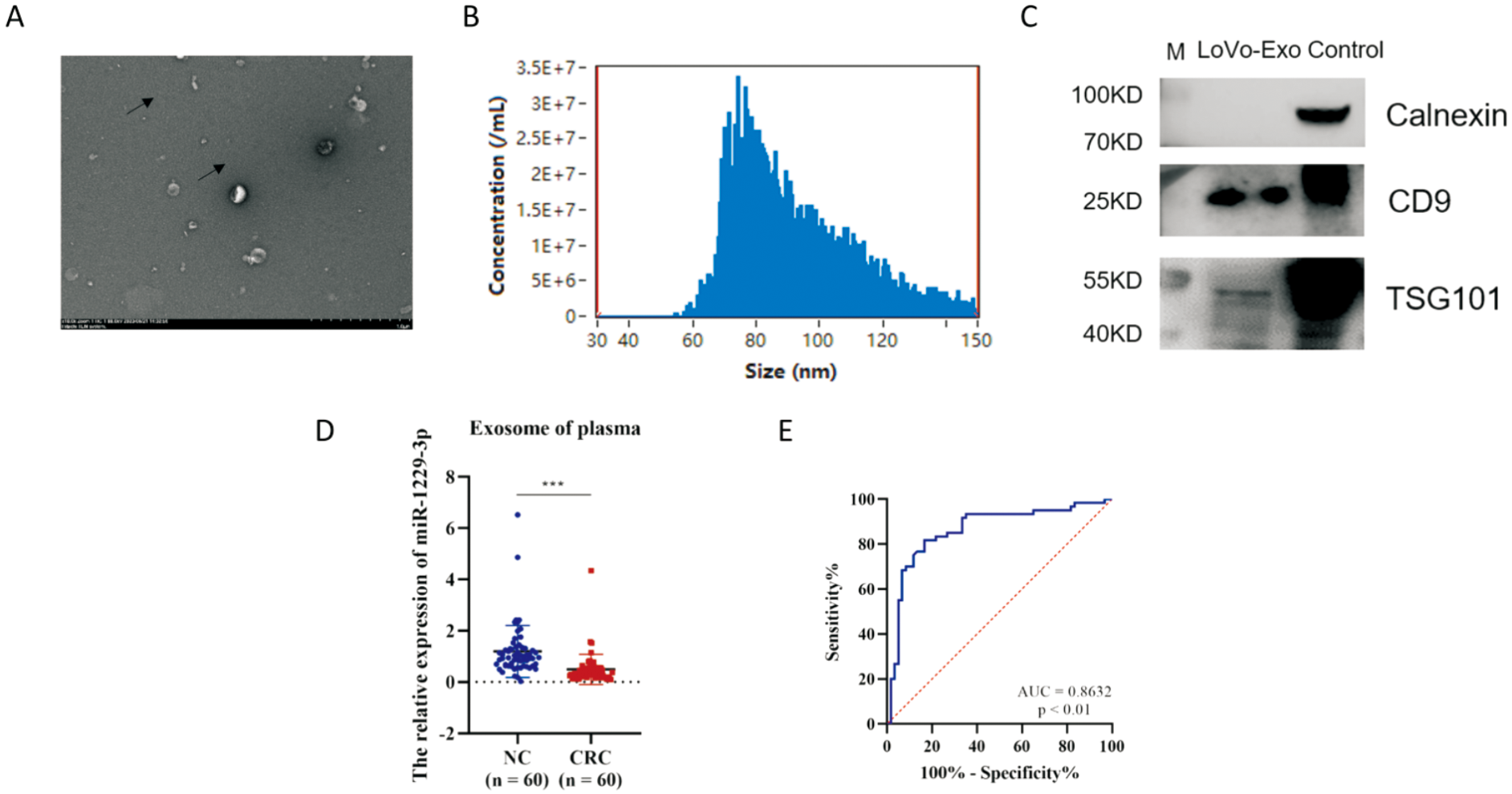
miR-1229-3p from plasma exosomes can distinguish patients with CRC A: Representative image of exosomes from LoVo cells under electron microscopy with a bar of 1.0 μm; B: Particle size analysis of exosomes from LoVo cells; C: Western blot analysis of exosomes positive markers TSG101, CD9 and negative marker Calnexin from LoVo cell exosomes, M is protein Marker, control is HEPG2 cells-CD9/Calnexin/TSG101; D: Expression of miR-1229-3p from plasma exosomes of 60 CRC patients and 60 healthy people; E: Receiver operating characteristic curve of miR-1229-3p on the diagnostic efficacy of CRC. ***:P<0.001.
| [1] |
SIEGEL R L, MILLER K D, WAGLE N S, et al. Cancer statistics, 2023[J]. CA Cancer J Clin, 2023, 73(1):17-48.
doi: 10.3322/caac.v73.1 URL |
| [2] | 中华人民共和国国家卫生健康委员会. 中国结直肠癌诊疗规范(2023版)[J]. 中华消化外科杂志, 2023, 22(6):667-698. |
| National Health Commission of the People′s Republic of China. Chinese protocol of diagnosis and treatment of colorectal cancer (2023 edition)[J]. Chin J Dig Surg, 2023, 22(6):667-698. | |
| [3] |
SUNG H, FERLAY J, SIEGEL R L, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries[J]. CA Cancer J Clin, 2021, 71(3):209-249.
doi: 10.3322/caac.v71.3 URL |
| [4] |
ALLEMANI C, MATSUDA T, DI CARLO V, et al. Global surveillance of trends in cancer survival 2000-14 (CONCORD-3): analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries[J]. Lancet, 2018, 391(10125):1023-1075.
doi: S0140-6736(17)33326-3 pmid: 29395269 |
| [5] | 张忠涛. 中国结直肠外科20年回顾与展望[J]. 中华消化外科杂志, 2022, 21(01):53-56. |
| ZHANG Z T. Two-decade retrospect and prospect of colorectal surgery in China[J]. Chin J Dig Surg, 2022, 21(1):53-56. | |
| [6] | 杨盈, 孟文建, 王自强. 结直肠癌的综合治疗[J]. 中华消化外科杂志, 2022, 21(06):753-765. |
| YANG Ying, MENG Wenjian, WANG Ziqiang. Comprehensive treatment of colorectal cancer[J]. Chin J Dig Surg, 2022, 21(06):753-765. | |
| [7] |
HARRIS T J, MCCORMICK F. The molecular pathology of cancer[J]. Nat Rev Clin Oncol, 2010, 7(5):251-265.
doi: 10.1038/nrclinonc.2010.41 pmid: 20351699 |
| [8] |
AMBROS V, BARTEL B, BARTEL D P, et al. A uniform system for microRNA annotation[J]. RNA, 2003, 9(3):277-279.
pmid: 12592000 |
| [9] |
POY M N, ELIASSON L, KRUTZFELDT J, et al. A pancreatic islet-specific microRNA regulates insulin secretion[J]. Nature, 2004, 432(7014):226-230.
doi: 10.1038/nature03076 |
| [10] |
CALAME K. MicroRNA-155 function in B Cells[J]. Immunity, 2007, 27(6):825-827.
doi: 10.1016/j.immuni.2007.11.010 pmid: 18093533 |
| [11] |
GRECO S J, RAMESHWAR P. MicroRNAs regulate synthesis of the neurotransmitter substance P in human me-senchymal stem cell-derived neuronal cells[J]. Proc Natl Acad Sci U S A, 2007, 104(39):15484-15489.
doi: 10.1073/pnas.0703037104 URL |
| [12] |
STEPONAITIENE R, KUPCINSKAS J, LANGNER C, et al. Epigenetic silencing of miR-137 is a frequent event in gastric carcinogenesis[J]. Mol Carcinog, 2016, 55(4):376-386.
doi: 10.1002/mc.22287 URL |
| [13] |
WANG F, YING H Q, HE B S, et al. Upregulated lncRNA-UCA1 contributes to progression of hepatocellular carcinoma through inhibition of miR-216b and activation of FGFR1/ERK signaling pathway[J]. Oncotarget, 2015, 6(10):7899-7917.
doi: 10.18632/oncotarget.3219 pmid: 25760077 |
| [14] |
LIU R, GU J, JIANG P, et al. DNMT1-microRNA126 epigenetic circuit contributes to esophageal squamous cell carcinoma growth via ADAM9-EGFR-AKT signaling[J]. Clin Cancer Res, 2015, 21(4):854-863.
doi: 10.1158/1078-0432.CCR-14-1740 pmid: 25512445 |
| [15] |
LEE Y S, DUTTA A. MicroRNAs in cancer[J]. Annu Rev Pathol, 2009, 4:199-227.
doi: 10.1146/annurev.pathol.4.110807.092222 pmid: 18817506 |
| [16] |
GILLIES J K, LORIMER I A. Regulation of p27Kip1 by miRNA 221/222 in glioblastoma[J]. Cell Cycle, 2007, 6(16):2005-2009.
doi: 10.4161/cc.6.16.4526 pmid: 17721077 |
| [17] |
JOHNSON C D, ESQUELA-KERSCHER A, STEFANI G, et al. The let-7 microRNA represses cell proliferation pathways in human cells[J]. Cancer Res, 2007, 67(16):7713-7722.
doi: 10.1158/0008-5472.CAN-07-1083 pmid: 17699775 |
| [18] |
MOTT J L, KOBAYASHI S, BRONK S F, et al. mir-29 regulates Mcl-1 protein expression and apoptosis[J]. Oncogene, 2007, 26(42):6133-6140.
pmid: 17404574 |
| [19] |
BOMMER G T, GERIN I, FENG Y, et al. p53-mediated activation of miRNA34 candidate tumor-suppressor genes[J]. Curr Biol, 2007, 17(15):1298-1307.
doi: 10.1016/j.cub.2007.06.068 pmid: 17656095 |
| [20] |
SCOTT G K, GOGA A, BHAUMIK D, et al. Coordinate suppression of ERBB2 and ERBB3 by enforced expression of micro-RNA miR-125a or miR-125b[J]. J Biol Chem, 2007, 282(2):1479-1486.
doi: 10.1074/jbc.M609383200 pmid: 17110380 |
| [21] |
KUMAR M S, LU J, MERCER K L, et al. Impaired microRNA processing enhances cellular transformation and tumorigenesis[J]. Nat Genet, 2007, 39(5):673-677.
doi: 10.1038/ng2003 pmid: 17401365 |
| [22] |
JUNG G, HERNÁNDEZ-ILLÁN E, MOREIRA L, et al. Epigenetics of colorectal cancer: biomarker and therapeutic potential[J]. Nat Rev Gastroenterol Hepatol, 2020, 17(2):111-130.
doi: 10.1038/s41575-019-0230-y pmid: 31900466 |
| [23] |
FARHAN M, WANG H, GAUR U, et al. FOXO signaling pathways as therapeutic targets in cancer[J]. Int J Biol Sci, 2017, 13(7):815-827.
doi: 10.7150/ijbs.20052 pmid: 28808415 |
| [24] |
OUDHOFF M J, BRAAM M J S, FREEMAN S A, et al. SETD7 controls intestinal regeneration and tumorigene-sis by regulating Wnt/β-Catenin and Hippo/YAP signa-ling[J]. Dev Cell, 2016, 37(1):47-57.
doi: 10.1016/j.devcel.2016.03.002 URL |
| [25] |
DEWS M, HOMAYOUNI A, YU D, et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster[J]. Nat Genet, 2006, 38(9):1060-1065.
doi: 10.1038/ng1855 pmid: 16878133 |
| [26] |
MERTENS-TALCOTT S U, CHINTHARLAPALLI S, LI X, et al. The oncogenic microRNA-27a targets genes that regulate specificity protein transcription factors and the G2-M checkpoint in MDA-MB-231 breast cancer cells[J]. Cancer Res, 2007, 67(22):11001-110011.
doi: 10.1158/0008-5472.CAN-07-2416 URL |
| [27] |
HOU Y, ZHANG Q, PANG W, et al. YTHDC1-mediated augmentation of miR-30d in repressing pancreatic tumorigenesis via attenuation of RUNX1-induced transcriptional activation of Warburg effect[J]. Cell Death Differ, 2021, 28(11):3105-3124.
doi: 10.1038/s41418-021-00804-0 pmid: 34021267 |
| [28] |
KONOSHENKO M Y, BRYZGUNOVA O E, LAKTIONOV P P. miRNAs and androgen deprivation therapy for prostate cancer[J]. Biochim Biophys Acta Rev Cancer, 2021, 1876(2):188625.
doi: 10.1016/j.bbcan.2021.188625 URL |
| [29] |
HU X, CHEN Q, GUO H, et al. Identification of target PTEN-based miR-425 and miR-576 as potential diagnostic and immunotherapeutic biomarkers of colorectal cancer with liver metastasis[J]. Front Oncol, 2021, 11:657984.
doi: 10.3389/fonc.2021.657984 URL |
| [30] |
LONE W, BOUSKA A, SHARMA S, et al. Genome-wide miRNA expression profiling of molecular subgroups of peripheral T-cell lymphoma[J]. Clin Cancer Res, 2021, 27(21):6039-6053.
doi: 10.1158/1078-0432.CCR-21-0573 URL |
| [31] |
KONISHI H, SATO H, TAKAHASHI K, et al. Tumor-progressive mechanisms mediating miRNA-protein interaction[J]. Int J Mol Sci, 2021, 22(22):12303.
doi: 10.3390/ijms222212303 URL |
| [32] |
BUTKYTĖ S, ČIUPAS L, JAKUBAUSKIENĖ E, et al. Splicing-dependent expression of microRNAs of mirtron origin in human digestive and excretory system cancer cells[J]. Clin Epigenetics, 2016, 8:33.
doi: 10.1186/s13148-016-0200-y pmid: 27019673 |
| [33] |
NISHIBEPPU K, KOMATSU S, IMAMURA T, et al. Plasma microRNA profiles: identification of miR-1229-3p as a novel chemoresistant and prognostic biomarker in gastric cancer[J]. Sci Rep, 2020, 10(1):3161.
doi: 10.1038/s41598-020-59939-8 pmid: 32081926 |
| [34] |
ZHAO X, CUI L. A robust six-miRNA prognostic signature for head and neck squamous cell carcinoma[J]. J Cell Physiol, 2020, 235(11):8799-8811.
doi: 10.1002/jcp.29723 pmid: 32342519 |
| [35] |
ZHANG C, ZHANG Q, LI H, et al. miR-1229-3p as a prognostic predictor facilitates cell viability, migration, and invasion of hepatocellular carcinoma[J]. Horm Metab Res, 2021, 53(11):759-766.
doi: 10.1055/a-1646-8415 pmid: 34740278 |
| [36] | CAO L, WANG M, XU K. Research progress of role and mechanism of SETD7 in tumor occurrence and progression[J]. Zhongguo Fei Ai Za Zhi, 2023, 26(1):38-45. |
| [1] | WANG Shukui, GU Xinliang. Advances in the study of tsRNA as diagnostic and prognostic biomarkers in cancer [J]. Journal of Diagnostics Concepts & Practice, 2023, 22(05): 413-420. |
| [2] | CHEN Guoqun, CAI Jiaodi. Interpretation of the Clinical Practice Guidelines for Non-small Lung Cancer (version 4 and version 5) of 2022 National Comprehensive Cancer Nerwork(NCCN) [J]. Journal of Diagnostics Concepts & Practice, 2023, 22(01): 8-13. |
| [3] | WANG Han, LU Haidi, WANG Lei, CONG Wenming, ZHENG Jianming, BAI Chenguang. Clinicopathological features of 2 cases of squamous cell carcinoma and 2 cases of adenosquamous carcinoma [J]. Journal of Diagnostics Concepts & Practice, 2023, 22(01): 44-49. |
| [4] | LI Jiaxi, WANG Jinjiang, YU Liping, YUAN Ying, QIAO Guanglei, MA Lijun. Effect of RAB25 knockdown on ferroptosis of colorectal cancer cells [J]. Journal of Diagnostics Concepts & Practice, 2022, 21(06): 710-718. |
| [5] | WU Dongdong, CHEN Yuhui, LIU Fang, LIU Yinhong, JIANG Jingwen. Study progress on cerebral small vessel disease complicated with neurodegenerative disorders in central nervous system [J]. Journal of Diagnostics Concepts & Practice, 2022, 21(05): 644-649. |
| [6] | YANG Ruixin, DU Yutong, YAN Ranlin, ZHU Zhenggang, LI Chen, YU Yingyan. Improving exploration of biological sample pretreatment in single-cell transcriptome sequencing of gastrointestinal tumors [J]. Journal of Diagnostics Concepts & Practice, 2022, 21(05): 567-574. |
| [7] | ZHOU Sifeng, XU Haishu, FAN Xinsheng. Application of metabolomics of different biological samples in study of OSAHS biomarkers [J]. Journal of Diagnostics Concepts & Practice, 2022, 21(04): 535-540. |
| [8] | ZHANG Hua, LU Wei, YANG Chengyi, XIANG Mingjie. Value of serum FBLN1 detection in diagnosis and prognosis prediction of colorectal cancer [J]. Journal of Diagnostics Concepts & Practice, 2021, 20(05): 462-465. |
| [9] | XU Chenying, XU Qingling, TANG Chenyue, YU Lifen. Detection rate of colorectal adenoma in the asymptomatic population of 40 to 59 years and its relationship with the detected gastric polyps in Shanghai [J]. Journal of Diagnostics Concepts & Practice, 2020, 19(05): 504-509. |
| [10] | WANG Ziyuan, LIANG Tingyu, CHEN Jia, HAN Zhihong, WU Lili. Analysis of KRAS, NRAS and BRAF mutations in colorectal cancer and their correlation with clinicopathologic features and p53 protein expression [J]. Journal of Diagnostics Concepts & Practice, 2018, 17(06): 687-693. |
| [11] | YAO Jiejie, ZHU Ying, ZHAN Weiwei, CHEN Xiaosong, FEI Xiaochun. Correlation of sonographic features of non-mass ductal carcinoma in situ with clinicopathological findings and biomarkers [J]. Journal of Diagnostics Concepts & Practice, 2018, 17(06): 676-681. |
| [12] | DU Kun, YANG Xi, BIAN Binxian, REN Yiqian, ZHANG Guanghui. Comparison of diagnostic value of new infection biomarker presepsin with procalcitonin, C-reactive protein and interleukin-6 in diagnosis of bacterial infection [J]. Journal of Diagnostics Concepts & Practice, 2018, 17(05): 581-585. |
| [13] | CHEN Ying, HE Lili, JIANG Feng. Expression of long non-coding RNA FENDRR in human colorectal cancer and its correlation with prognosis [J]. Journal of Diagnostics Concepts & Practice, 2018, 17(01): 87-91. |
| [14] | . [J]. Journal of Diagnostics Concepts & Practice, 2015, 14(05): 455-460. |
| [15] | . [J]. Journal of Diagnostics Concepts & Practice, 2014, 13(06): 588-592. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||